from pathlib import Path
import os
import warnings
os.environ.setdefault("OMP_NUM_THREADS", "1")
os.environ.setdefault("OPENBLAS_NUM_THREADS", "1")
os.environ.setdefault("MKL_NUM_THREADS", "1")
os.environ.setdefault("NUMEXPR_NUM_THREADS", "1")
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import display
from sklearn.base import clone
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_absolute_error, mean_squared_error, r2_score
from sklearn.model_selection import GroupKFold
from sklearn.preprocessing import StandardScaler
from lightgbm import LGBMRegressor
from xgboost import XGBRegressor
warnings.filterwarnings("ignore", category=FutureWarning)
sns.set_theme(style="whitegrid", context="notebook")
plt.rcParams["figure.figsize"] = (11, 6)
plt.rcParams["axes.titlesize"] = 13
plt.rcParams["axes.labelsize"] = 11
pd.set_option("display.max_columns", 180)
pd.set_option("display.max_colwidth", 140)
PROJECT_ROOT = Path.cwd().resolve()
while not (PROJECT_ROOT / "data").exists() and PROJECT_ROOT.parent != PROJECT_ROOT:
PROJECT_ROOT = PROJECT_ROOT.parent
PROCESSED_DIR = PROJECT_ROOT / "data" / "processed"
NOTEBOOK_DIR = PROJECT_ROOT / "notebooks" / "discovery_quality_mediation"
WRITEUP_DIR = NOTEBOOK_DIR / "writeup"
FIGURE_DIR = WRITEUP_DIR / "figures"
TABLE_DIR = WRITEUP_DIR / "tables"
FIGURE_DIR.mkdir(parents=True, exist_ok=True)
TABLE_DIR.mkdir(parents=True, exist_ok=True)
ANALYSIS_PANEL_INPUT = PROCESSED_DIR / "kuairec_discovery_quality_mediation_analysis_panel.parquet"
EFFECT_SUMMARY_INPUT = PROCESSED_DIR / "kuairec_discovery_quality_effect_summary.csv"
ROBUSTNESS_SUMMARY_INPUT = PROCESSED_DIR / "kuairec_discovery_quality_robustness_summary.csv"
SEM_PATH_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_sem_path_results.csv"
SEM_BOOTSTRAP_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_sem_bootstrap.csv"
ML_EFFECTS_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_advanced_ml_effects.csv"
ML_PERFORMANCE_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_advanced_ml_performance.csv"
HETEROGENEITY_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_advanced_heterogeneity.csv"
FEATURE_IMPORTANCE_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_advanced_feature_importance.csv"
ADVANCED_SUMMARY_OUTPUT = PROCESSED_DIR / "kuairec_discovery_quality_advanced_model_summary.csv"Advanced SEM and ML Mediation Models
The previous notebooks built the discovery-quality mediation design, estimated a transparent linear g-computation decomposition, and stress-tested the result across definitions and specifications. This notebook adds a more advanced modeling layer.
The goal is not to replace the earlier notebooks. The goal is to ask whether the same story survives when we use more flexible models and a more formal path-model framing.
This notebook has four advanced pieces:
- SEM-style path model: estimate the
A -> M,M -> Y, andA -> Ypaths as a formal mediation path decomposition. - Cross-fitted ML nuisance models: estimate mediator and outcome nuisance functions with Linear Regression, LightGBM, and XGBoost without evaluating each row on models trained on that row.
- Heterogeneity analysis: check whether the discovery effect differs across prior activity, prior satisfaction, prior discovery, and user activity groups.
- Feature importance audit: inspect which covariates the tree-based nuisance models use most.
The main question remains the same: does high discovery exposure increase future user value, and is that effect meaningfully routed through same-day satisfaction depth?
1. Load Libraries and Paths
This cell imports the modeling libraries used in the notebook. LightGBM and XGBoost are used for flexible nuisance models, while linear models provide the transparent baseline and SEM-style path estimates. The path setup mirrors earlier notebooks so the outputs land in the same processed and writeup folders.
The thread settings keep LightGBM and XGBoost from taking over the machine. The dataset is small enough that single-threaded tree models are still fast and easier to reproduce.
2. Load Analysis Inputs
This cell loads the saved mediation analysis panel and the previous result tables. The advanced notebook should not reconstruct the panel from raw logs; it should reuse the cleaned analysis object created earlier.
analysis_panel = pd.read_parquet(ANALYSIS_PANEL_INPUT)
effect_summary = pd.read_csv(EFFECT_SUMMARY_INPUT)
robustness_summary = pd.read_csv(ROBUSTNESS_SUMMARY_INPUT)
load_summary = pd.DataFrame(
{
"artifact": ["analysis_panel", "effect_summary", "robustness_summary"],
"rows": [len(analysis_panel), len(effect_summary), len(robustness_summary)],
"columns": [analysis_panel.shape[1], effect_summary.shape[1], robustness_summary.shape[1]],
}
)
main_effects = effect_summary.query(
"outcome == 'Y_future_interactions' and estimand in ['gcomp_total_effect', 'natural_direct_effect', 'natural_indirect_effect']"
)[["estimand", "estimate", "ci_95_lower", "ci_95_upper", "relative_effect"]]
display(load_summary)
display(main_effects.round(4))
display(robustness_summary.round(4))| artifact | rows | columns | |
|---|---|---|---|
| 0 | analysis_panel | 8199 | 103 |
| 1 | effect_summary | 21 | 12 |
| 2 | robustness_summary | 4 | 9 |
| estimand | estimate | ci_95_lower | ci_95_upper | relative_effect | |
|---|---|---|---|---|---|
| 2 | gcomp_total_effect | 37.2781 | 32.2444 | 42.8135 | 0.1152 |
| 3 | natural_direct_effect | 38.5858 | 33.3634 | 43.7154 | 0.1192 |
| 4 | natural_indirect_effect | -1.3077 | -2.3047 | -0.4290 | -0.0036 |
| robustness_family | specifications | total_effect_min | total_effect_max | total_effect_share_positive | indirect_effect_min | indirect_effect_max | indirect_effect_share_positive | median_abs_indirect_to_total_ratio | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | threshold_sensitivity | 4 | 37.2781 | 40.0555 | 1.0 | -1.6532 | -1.3077 | 0.0 | 0.0402 |
| 1 | mediator_sensitivity | 5 | 37.2560 | 37.3502 | 1.0 | -1.7145 | 0.1281 | 0.2 | 0.0359 |
| 2 | model_sensitivity | 5 | 34.8085 | 37.6289 | 1.0 | -1.5581 | -0.9360 | 0.0 | 0.0351 |
| 3 | outcome_sensitivity | 1 | 37.2781 | 37.2781 | 1.0 | -1.3077 | -1.3077 | 0.0 | 0.0351 |
The earlier result is the benchmark. The advanced models should be judged against this reference, not treated as automatically better just because they are more flexible.
3. Define Variables and Control Sets
This cell defines the treatment, mediator, outcome, and covariates. The covariates are the same pre-treatment history and profile features used in prior notebooks. This keeps the comparison focused on model form rather than changing the adjustment set.
TREATMENT = "A_high_discovery"
MEDIATOR = "M_satisfaction_depth"
PRIMARY_OUTCOME = "Y_future_interactions"
SECONDARY_OUTCOME = "Y_future_play_hours"
WEIGHT_COLUMN = "stabilized_ipw_capped"
GROUP_COLUMN = "user_id"
RANDOM_SEED = 42
N_SPLITS = 3
SEM_BOOTSTRAP_ITERATIONS = 100
base_numeric_covariates = [
"calendar_day_index",
"lag_1_active_day",
"prior_3day_active_day",
"lag_1_interactions",
"prior_3day_interactions",
"lag_1_total_play_duration_sec",
"prior_3day_total_play_duration_sec",
"lag_1_valid_play_share",
"prior_3day_valid_play_share",
"lag_1_high_satisfaction_share",
"prior_3day_high_satisfaction_share",
"lag_1_discovery_candidate_share",
"prior_3day_discovery_candidate_share",
"recent_activity_score",
"register_days",
"follow_user_num",
"fans_user_num",
"friend_user_num",
"is_lowactive_period",
"is_live_streamer",
"is_video_author",
]
profile_onehot_covariates = [col for col in analysis_panel.columns if col.startswith("onehot_feat")]
numeric_covariates = [col for col in base_numeric_covariates + profile_onehot_covariates if col in analysis_panel.columns]
categorical_covariates = [
col
for col in [
"user_active_degree",
"follow_user_num_range",
"fans_user_num_range",
"friend_user_num_range",
"register_days_range",
]
if col in analysis_panel.columns
]
variable_summary = pd.DataFrame(
{
"item": ["treatment", "mediator", "primary_outcome", "secondary_outcome", "numeric_covariates", "categorical_covariates", "group_folds"],
"value": [TREATMENT, MEDIATOR, PRIMARY_OUTCOME, SECONDARY_OUTCOME, len(numeric_covariates), len(categorical_covariates), N_SPLITS],
}
)
display(variable_summary)| item | value | |
|---|---|---|
| 0 | treatment | A_high_discovery |
| 1 | mediator | M_satisfaction_depth |
| 2 | primary_outcome | Y_future_interactions |
| 3 | secondary_outcome | Y_future_play_hours |
| 4 | numeric_covariates | 39 |
| 5 | categorical_covariates | 5 |
| 6 | group_folds | 3 |
The cross-fitting group is user_id, so the same user’s days do not appear in both train and validation folds. That is important because each user contributes repeated active days.
4. Build the Modeling Matrix
This cell prepares the adjustment matrix used by the SEM and ML models. Numeric columns are median-filled, categorical columns are one-hot encoded, and constant columns are removed.
def make_safe_column_names(columns):
safe_names = []
counts = {}
for col in columns:
safe = "".join(ch if ch.isalnum() else "_" for ch in str(col))
safe = "_".join(part for part in safe.split("_") if part)
if not safe:
safe = "feature"
if safe[0].isdigit():
safe = f"feature_{safe}"
counts[safe] = counts.get(safe, 0) + 1
if counts[safe] > 1:
safe = f"{safe}_{counts[safe]}"
safe_names.append(safe)
return safe_names
def build_covariate_matrix(frame, numeric_cols, categorical_cols):
numeric = frame[numeric_cols].copy()
for col in numeric.columns:
numeric[col] = pd.to_numeric(numeric[col], errors="coerce")
numeric[col] = numeric[col].fillna(numeric[col].median())
if categorical_cols:
categorical = frame[categorical_cols].copy()
for col in categorical.columns:
categorical[col] = categorical[col].astype("string").fillna("missing")
encoded = pd.get_dummies(categorical, drop_first=True, dtype=float)
design = pd.concat([numeric, encoded], axis=1)
else:
design = numeric
design = design.loc[:, design.nunique(dropna=False) > 1]
design.columns = make_safe_column_names(design.columns)
return design.astype(float)
X_covariates = build_covariate_matrix(analysis_panel, numeric_covariates, categorical_covariates)
A = analysis_panel[TREATMENT].astype(float).to_numpy()
M = analysis_panel[MEDIATOR].astype(float).to_numpy()
Y = analysis_panel[PRIMARY_OUTCOME].astype(float).to_numpy()
weights = analysis_panel[WEIGHT_COLUMN].astype(float).to_numpy()
groups = analysis_panel[GROUP_COLUMN].to_numpy()
matrix_summary = pd.DataFrame(
{
"item": ["rows", "covariate_columns", "treatment_rate", "mediator_mean", "outcome_mean", "unique_users"],
"value": [len(analysis_panel), X_covariates.shape[1], A.mean(), M.mean(), Y.mean(), analysis_panel[GROUP_COLUMN].nunique()],
}
)
display(matrix_summary)
display(X_covariates.head())| item | value | |
|---|---|---|
| 0 | rows | 8199.000000 |
| 1 | covariate_columns | 54.000000 |
| 2 | treatment_rate | 0.500061 |
| 3 | mediator_mean | 0.618626 |
| 4 | outcome_mean | 340.694475 |
| 5 | unique_users | 133.000000 |
| calendar_day_index | lag_1_active_day | prior_3day_active_day | lag_1_interactions | prior_3day_interactions | lag_1_total_play_duration_sec | prior_3day_total_play_duration_sec | lag_1_valid_play_share | prior_3day_valid_play_share | lag_1_high_satisfaction_share | prior_3day_high_satisfaction_share | lag_1_discovery_candidate_share | prior_3day_discovery_candidate_share | recent_activity_score | register_days | follow_user_num | fans_user_num | friend_user_num | is_video_author | onehot_feat0 | onehot_feat1 | onehot_feat2 | onehot_feat3 | onehot_feat4 | onehot_feat6 | onehot_feat7 | onehot_feat8 | onehot_feat9 | onehot_feat10 | onehot_feat11 | onehot_feat12 | onehot_feat13 | onehot_feat14 | onehot_feat15 | onehot_feat17 | user_active_degree_full_active | user_active_degree_high_active | follow_user_num_range_10_50 | follow_user_num_range_100_150 | follow_user_num_range_150_250 | follow_user_num_range_250_500 | follow_user_num_range_50_100 | follow_user_num_range_0 | follow_user_num_range_500 | fans_user_num_range_1_10 | fans_user_num_range_10_100 | friend_user_num_range_1_5 | friend_user_num_range_30_60 | friend_user_num_range_5_30 | register_days_range_31_60 | register_days_range_366_730 | register_days_range_61_90 | register_days_range_730 | register_days_range_91_180 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.000 | 0.000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 224.0 | 7.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 24.0 | 876.0 | 1.0 | 1.0 | 4.0 | 98.0 | 6.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 1 | 1.0 | 1.0 | 1.0 | 32.0 | 32.0 | 163.970 | 163.970 | 0.937500 | 0.937500 | 0.156250 | 0.156250 | 0.687500 | 0.687500 | 0.562990 | 224.0 | 7.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 24.0 | 876.0 | 1.0 | 1.0 | 4.0 | 98.0 | 6.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 2 | 2.0 | 1.0 | 2.0 | 20.0 | 52.0 | 130.986 | 294.956 | 0.950000 | 1.887500 | 0.350000 | 0.506250 | 0.450000 | 1.137500 | 0.639277 | 224.0 | 7.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 24.0 | 876.0 | 1.0 | 1.0 | 4.0 | 98.0 | 6.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 3 | 3.0 | 1.0 | 3.0 | 16.0 | 68.0 | 100.920 | 395.876 | 1.000000 | 2.887500 | 0.187500 | 0.693750 | 0.437500 | 1.575000 | 0.681755 | 224.0 | 7.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 24.0 | 876.0 | 1.0 | 1.0 | 4.0 | 98.0 | 6.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 4 | 4.0 | 1.0 | 3.0 | 37.0 | 73.0 | 222.720 | 454.626 | 0.891892 | 2.841892 | 0.216216 | 0.753716 | 0.432432 | 1.319932 | 0.693019 | 224.0 | 7.0 | 3.0 | 0.0 | 0.0 | 0.0 | 1.0 | 24.0 | 876.0 | 1.0 | 1.0 | 4.0 | 98.0 | 6.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
The modeling matrix is intentionally reused across methods. SEM, Linear Regression, LightGBM, and XGBoost all see the same observed adjustment information.
5. SEM-Style Path Model
This cell estimates a simple path model:
M = alpha * A + X + errorY = c_prime * A + beta * M + X + errorindirect = alpha * betatotal_path = c_prime + alpha * beta
This is not a full latent-variable SEM. It is a transparent path-analysis version of the mediation story, which is often enough for product analytics settings where the variables are observed scores.
def design_with_columns(covariates, **named_arrays):
design = covariates.copy()
for name, values in reversed(list(named_arrays.items())):
design.insert(0, name, np.asarray(values, dtype=float))
return design
def fit_sem_path(frame, covariates):
a = frame[TREATMENT].astype(float).to_numpy()
m = frame[MEDIATOR].astype(float).to_numpy()
y = frame[PRIMARY_OUTCOME].astype(float).to_numpy()
w = frame[WEIGHT_COLUMN].astype(float).to_numpy()
mediator_design = design_with_columns(covariates, A=a)
mediator_model = LinearRegression().fit(mediator_design, m, sample_weight=w)
alpha = float(mediator_model.coef_[0])
outcome_design = design_with_columns(covariates, A=a, M=m)
outcome_model = LinearRegression().fit(outcome_design, y, sample_weight=w)
c_prime = float(outcome_model.coef_[0])
beta = float(outcome_model.coef_[1])
total_design = design_with_columns(covariates, A=a)
total_model = LinearRegression().fit(total_design, y, sample_weight=w)
total_coefficient = float(total_model.coef_[0])
return {
"alpha_A_to_M": alpha,
"beta_M_to_Y": beta,
"c_prime_A_to_Y": c_prime,
"indirect_alpha_beta": alpha * beta,
"total_path_c_prime_plus_indirect": c_prime + alpha * beta,
"total_regression_coefficient": total_coefficient,
"mediator_model_r2": mediator_model.score(mediator_design, m, sample_weight=w),
"outcome_model_r2": outcome_model.score(outcome_design, y, sample_weight=w),
"total_model_r2": total_model.score(total_design, y, sample_weight=w),
}
sem_path_results = pd.DataFrame([fit_sem_path(analysis_panel, X_covariates)])
display(sem_path_results.round(4))| alpha_A_to_M | beta_M_to_Y | c_prime_A_to_Y | indirect_alpha_beta | total_path_c_prime_plus_indirect | total_regression_coefficient | mediator_model_r2 | outcome_model_r2 | total_model_r2 | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.0231 | -48.4626 | 38.4311 | -1.1172 | 37.3139 | 37.3139 | 0.4653 | 0.6642 | 0.6637 |
The SEM-style table gives a compact path decomposition. If alpha is positive but beta is negative, high discovery can increase satisfaction depth while satisfaction depth is not the channel that raises future interaction count.
6. Bootstrap the SEM-Style Paths by User
This cell bootstraps the path model by resampling users. The purpose is to quantify how stable the SEM-style path coefficients are under the repeated-user structure.
rng = np.random.default_rng(RANDOM_SEED)
unique_users = np.sort(analysis_panel[GROUP_COLUMN].unique())
indexed_panel = analysis_panel.set_index(GROUP_COLUMN, drop=False)
sem_bootstrap_rows = []
for iteration in range(SEM_BOOTSTRAP_ITERATIONS):
sampled_users = rng.choice(unique_users, size=len(unique_users), replace=True)
sample = indexed_panel.loc[sampled_users].reset_index(drop=True)
sample_covariates = build_covariate_matrix(sample, numeric_covariates, categorical_covariates)
sample_covariates = sample_covariates.reindex(columns=X_covariates.columns, fill_value=0.0)
row = fit_sem_path(sample, sample_covariates)
row["bootstrap_iteration"] = iteration
sem_bootstrap_rows.append(row)
sem_bootstrap = pd.DataFrame(sem_bootstrap_rows)
sem_interval_rows = []
for column in [
"alpha_A_to_M",
"beta_M_to_Y",
"c_prime_A_to_Y",
"indirect_alpha_beta",
"total_path_c_prime_plus_indirect",
"total_regression_coefficient",
]:
sem_interval_rows.append(
{
"path_quantity": column,
"estimate": sem_path_results.loc[0, column],
"bootstrap_mean": sem_bootstrap[column].mean(),
"ci_95_lower": sem_bootstrap[column].quantile(0.025),
"ci_95_upper": sem_bootstrap[column].quantile(0.975),
}
)
sem_path_summary = pd.DataFrame(sem_interval_rows)
display(sem_path_summary.round(4))| path_quantity | estimate | bootstrap_mean | ci_95_lower | ci_95_upper | |
|---|---|---|---|---|---|
| 0 | alpha_A_to_M | 0.0231 | 0.0235 | 0.0183 | 0.0281 |
| 1 | beta_M_to_Y | -48.4626 | -48.4765 | -74.0799 | -21.9557 |
| 2 | c_prime_A_to_Y | 38.4311 | 38.7951 | 32.6143 | 44.0143 |
| 3 | indirect_alpha_beta | -1.1172 | -1.1509 | -1.8332 | -0.4317 |
| 4 | total_path_c_prime_plus_indirect | 37.3139 | 37.6442 | 31.4660 | 42.8712 |
| 5 | total_regression_coefficient | 37.3139 | 37.6442 | 31.4660 | 42.8712 |
The bootstrapped path table is the formal counterpart to the earlier g-computation results. It lets us compare whether the simple path-product indirect effect points in the same direction as the notebook 04 indirect estimate.
7. Plot SEM Path Quantities
This cell plots the SEM-style path quantities with bootstrap intervals. The visual focuses on the direct path, indirect path, and total path because these are the quantities that correspond most closely to the earlier decomposition.
sem_plot = sem_path_summary.query(
"path_quantity in ['c_prime_A_to_Y', 'indirect_alpha_beta', 'total_path_c_prime_plus_indirect', 'total_regression_coefficient']"
).copy()
sem_plot["label"] = sem_plot["path_quantity"].map(
{
"c_prime_A_to_Y": "Direct path",
"indirect_alpha_beta": "Indirect path",
"total_path_c_prime_plus_indirect": "Direct + indirect path",
"total_regression_coefficient": "Total regression path",
}
)
sem_plot["lower_error"] = sem_plot["estimate"] - sem_plot["ci_95_lower"]
sem_plot["upper_error"] = sem_plot["ci_95_upper"] - sem_plot["estimate"]
fig, ax = plt.subplots(figsize=(10, 5.5))
sns.barplot(data=sem_plot, x="estimate", y="label", color="steelblue", ax=ax)
for row_index, row in sem_plot.reset_index(drop=True).iterrows():
ax.errorbar(
x=row["estimate"],
y=row_index,
xerr=[[row["lower_error"]], [row["upper_error"]]],
fmt="none",
color="black",
capsize=4,
linewidth=1.2,
)
ax.axvline(0, color="black", linewidth=1)
ax.set_title("SEM-Style Path Decomposition")
ax.set_xlabel("Effect on future 7-day interactions")
ax.set_ylabel("")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "21_sem_path_decomposition.png", dpi=160, bbox_inches="tight")
plt.show()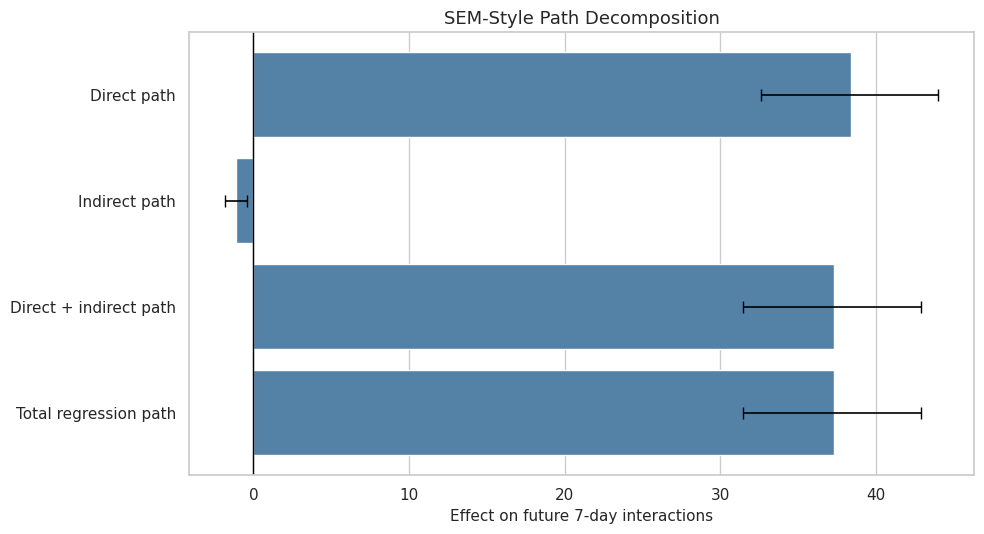
The SEM plot should be read as a path-model summary, not as a new identification strategy. It is useful because it compresses the mediation story into a few path coefficients.
8. Define Cross-Fitted ML Nuisance Models
This cell defines the model families used in the cross-fitted g-computation. The Linear model is the transparent baseline. LightGBM and XGBoost allow nonlinearities and interactions in the nuisance functions.
model_factories = {
"linear": lambda: LinearRegression(),
"lightgbm": lambda: LGBMRegressor(
n_estimators=80,
learning_rate=0.05,
num_leaves=24,
subsample=0.85,
colsample_bytree=0.85,
random_state=RANDOM_SEED,
n_jobs=1,
verbose=-1,
),
"xgboost": lambda: XGBRegressor(
n_estimators=80,
max_depth=3,
learning_rate=0.05,
subsample=0.85,
colsample_bytree=0.85,
objective="reg:squarederror",
tree_method="hist",
random_state=RANDOM_SEED,
n_jobs=1,
verbosity=0,
),
}
model_summary = pd.DataFrame(
{
"model_family": list(model_factories.keys()),
"role": [
"transparent benchmark",
"tree-based nonlinear nuisance model",
"boosted-tree nonlinear nuisance model",
],
}
)
display(model_summary)| model_family | role | |
|---|---|---|
| 0 | linear | transparent benchmark |
| 1 | lightgbm | tree-based nonlinear nuisance model |
| 2 | xgboost | boosted-tree nonlinear nuisance model |
All three model families estimate the same counterfactual quantities. That keeps the comparison fair: any differences come from nuisance model flexibility, not from changing the causal estimand.
9. Implement Cross-Fitted ML G-Computation
This cell implements group cross-fitting. For each fold, models are trained on some users and evaluated on held-out users. The estimator predicts M(0), M(1), Y(0, M(0)), Y(1, M(0)), and Y(1, M(1)) out of fold, then averages the implied effects.
def treatment_design(covariates, treatment_values):
design = covariates.copy()
design.insert(0, TREATMENT, np.asarray(treatment_values, dtype=float))
return design
def outcome_design(covariates, treatment_values, mediator_values):
treatment_array = np.asarray(treatment_values, dtype=float)
mediator_array = np.asarray(mediator_values, dtype=float)
design = covariates.copy()
design.insert(0, "A_x_M", treatment_array * mediator_array)
design.insert(0, MEDIATOR, mediator_array)
design.insert(0, TREATMENT, treatment_array)
return design
def fit_with_optional_weight(model, X, y, sample_weight=None):
try:
model.fit(X, y, sample_weight=sample_weight)
except TypeError:
model.fit(X, y)
return model
def regression_metrics(y_true, y_pred):
rmse = mean_squared_error(y_true, y_pred) ** 0.5
return {
"rmse": rmse,
"mae": mean_absolute_error(y_true, y_pred),
"r2": r2_score(y_true, y_pred),
}
def crossfit_ml_mediation(model_name, model_factory, outcome_col=PRIMARY_OUTCOME):
n = len(analysis_panel)
splitter = GroupKFold(n_splits=N_SPLITS)
m_hat = np.zeros(n)
y_hat = np.zeros(n)
m0 = np.zeros(n)
m1 = np.zeros(n)
y_0_m0 = np.zeros(n)
y_1_m0 = np.zeros(n)
y_1_m1 = np.zeros(n)
y_0_m1 = np.zeros(n)
fold_rows = []
for fold_id, (train_idx, test_idx) in enumerate(splitter.split(X_covariates, A, groups=groups), start=1):
X_train = X_covariates.iloc[train_idx]
X_test = X_covariates.iloc[test_idx]
A_train = A[train_idx]
M_train = M[train_idx]
Y_train = analysis_panel[outcome_col].astype(float).to_numpy()[train_idx]
W_train = weights[train_idx]
mediator_model = model_factory()
outcome_model = model_factory()
mediator_model = fit_with_optional_weight(
mediator_model,
treatment_design(X_train, A_train),
M_train,
sample_weight=W_train,
)
train_outcome_design = outcome_design(X_train, A_train, M_train)
outcome_model = fit_with_optional_weight(
outcome_model,
train_outcome_design,
Y_train,
sample_weight=W_train,
)
zeros = np.zeros(len(test_idx))
ones = np.ones(len(test_idx))
observed_a = A[test_idx]
observed_m = M[test_idx]
m_hat[test_idx] = mediator_model.predict(treatment_design(X_test, observed_a)).clip(0, 1)
m0[test_idx] = mediator_model.predict(treatment_design(X_test, zeros)).clip(0, 1)
m1[test_idx] = mediator_model.predict(treatment_design(X_test, ones)).clip(0, 1)
y_hat[test_idx] = outcome_model.predict(outcome_design(X_test, observed_a, observed_m))
y_0_m0[test_idx] = outcome_model.predict(outcome_design(X_test, zeros, m0[test_idx]))
y_1_m0[test_idx] = outcome_model.predict(outcome_design(X_test, ones, m0[test_idx]))
y_1_m1[test_idx] = outcome_model.predict(outcome_design(X_test, ones, m1[test_idx]))
y_0_m1[test_idx] = outcome_model.predict(outcome_design(X_test, zeros, m1[test_idx]))
fold_rows.append(
{
"model_family": model_name,
"outcome": outcome_col,
"fold": fold_id,
"train_rows": len(train_idx),
"test_rows": len(test_idx),
"test_users": len(np.unique(groups[test_idx])),
}
)
observed_y = analysis_panel[outcome_col].astype(float).to_numpy()
mediator_metrics = regression_metrics(M, m_hat)
outcome_metrics = regression_metrics(observed_y, y_hat)
effects = {
"model_family": model_name,
"outcome": outcome_col,
"outcome_label": "Future 7-day interactions" if outcome_col == PRIMARY_OUTCOME else "Future 7-day play hours",
"gcomp_total_effect": float(np.mean(y_1_m1 - y_0_m0)),
"natural_direct_effect": float(np.mean(y_1_m0 - y_0_m0)),
"natural_indirect_effect": float(np.mean(y_1_m1 - y_1_m0)),
"reverse_indirect_check": float(np.mean(y_0_m1 - y_0_m0)),
"mediator_shift": float(np.mean(m1 - m0)),
"reference_mean": float(np.mean(y_0_m0)),
"relative_total_effect": float(np.mean(y_1_m1 - y_0_m0) / np.mean(y_0_m0)),
}
performance = [
{"model_family": model_name, "outcome": outcome_col, "target": "mediator", **mediator_metrics},
{"model_family": model_name, "outcome": outcome_col, "target": "outcome", **outcome_metrics},
]
row_level = pd.DataFrame(
{
"user_id": analysis_panel[GROUP_COLUMN].to_numpy(),
"event_date": analysis_panel["event_date"].to_numpy(),
"model_family": model_name,
"outcome": outcome_col,
"ite_total": y_1_m1 - y_0_m0,
"ite_direct": y_1_m0 - y_0_m0,
"ite_indirect": y_1_m1 - y_1_m0,
"m0_hat": m0,
"m1_hat": m1,
"m_hat": m_hat,
"y_hat": y_hat,
}
)
return effects, performance, pd.DataFrame(fold_rows), row_level
print("Cross-fitted ML mediation function ready.")Cross-fitted ML mediation function ready.The row-level output is useful for heterogeneity. Each held-out row receives an estimated total, direct, and indirect effect, which can then be summarized by user segments.
10. Run Cross-Fitted ML Mediation
This cell runs the cross-fitted estimator for Linear Regression, LightGBM, and XGBoost. It estimates the primary future-interaction outcome and a secondary future-play-hours outcome.
ml_effect_rows = []
ml_performance_rows = []
fold_summaries = []
row_level_effect_frames = []
for outcome_col in [PRIMARY_OUTCOME, SECONDARY_OUTCOME]:
for model_name, model_factory in model_factories.items():
effects, performance, folds, row_level = crossfit_ml_mediation(
model_name,
model_factory,
outcome_col=outcome_col,
)
ml_effect_rows.append(effects)
ml_performance_rows.extend(performance)
fold_summaries.append(folds)
row_level_effect_frames.append(row_level)
ml_effects = pd.DataFrame(ml_effect_rows)
ml_performance = pd.DataFrame(ml_performance_rows)
ml_fold_summary = pd.concat(fold_summaries, ignore_index=True)
row_level_effects = pd.concat(row_level_effect_frames, ignore_index=True)
display(ml_effects.round(4))
display(ml_performance.round(4))| model_family | outcome | outcome_label | gcomp_total_effect | natural_direct_effect | natural_indirect_effect | reverse_indirect_check | mediator_shift | reference_mean | relative_total_effect | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | linear | Y_future_interactions | Future 7-day interactions | 37.3421 | 38.6464 | -1.3043 | -0.9904 | 0.0231 | 322.8968 | 0.1156 |
| 1 | lightgbm | Y_future_interactions | Future 7-day interactions | 3.2069 | 3.2652 | -0.0583 | -0.0137 | 0.0184 | 339.1834 | 0.0095 |
| 2 | xgboost | Y_future_interactions | Future 7-day interactions | 2.6829 | 2.7279 | -0.0450 | -0.0294 | 0.0138 | 339.7969 | 0.0079 |
| 3 | linear | Y_future_play_hours | Future 7-day play hours | 0.0786 | 0.0745 | 0.0042 | 0.0021 | 0.0231 | 0.7480 | 0.1051 |
| 4 | lightgbm | Y_future_play_hours | Future 7-day play hours | 0.0076 | 0.0030 | 0.0046 | 0.0046 | 0.0184 | 0.8177 | 0.0093 |
| 5 | xgboost | Y_future_play_hours | Future 7-day play hours | 0.0062 | 0.0034 | 0.0028 | 0.0027 | 0.0138 | 0.8230 | 0.0075 |
| model_family | outcome | target | rmse | mae | r2 | |
|---|---|---|---|---|---|---|
| 0 | linear | Y_future_interactions | mediator | 0.1015 | 0.0703 | 0.1878 |
| 1 | linear | Y_future_interactions | outcome | 96.6374 | 71.1050 | 0.7129 |
| 2 | lightgbm | Y_future_interactions | mediator | 0.0875 | 0.0629 | 0.3959 |
| 3 | lightgbm | Y_future_interactions | outcome | 66.7067 | 47.0941 | 0.8632 |
| 4 | xgboost | Y_future_interactions | mediator | 0.0870 | 0.0625 | 0.4026 |
| 5 | xgboost | Y_future_interactions | outcome | 66.0357 | 46.7289 | 0.8659 |
| 6 | linear | Y_future_play_hours | mediator | 0.1015 | 0.0703 | 0.1878 |
| 7 | linear | Y_future_play_hours | outcome | 0.3609 | 0.2428 | 0.4056 |
| 8 | lightgbm | Y_future_play_hours | mediator | 0.0875 | 0.0629 | 0.3959 |
| 9 | lightgbm | Y_future_play_hours | outcome | 0.2388 | 0.1672 | 0.7398 |
| 10 | xgboost | Y_future_play_hours | mediator | 0.0870 | 0.0625 | 0.4026 |
| 11 | xgboost | Y_future_play_hours | outcome | 0.2383 | 0.1694 | 0.7409 |
The model comparison is now cross-fitted. If the flexible models preserve the positive total effect and small mediated pathway, the earlier story is less likely to be an artifact of linear functional form.
11. Plot Cross-Fitted Model Comparison
This cell visualizes total, direct, and indirect effects across model families for the primary outcome. The plot uses the cross-fitted estimates from held-out users.
primary_ml_plot = ml_effects.query("outcome == @PRIMARY_OUTCOME").melt(
id_vars=["model_family"],
value_vars=["gcomp_total_effect", "natural_direct_effect", "natural_indirect_effect"],
var_name="effect_type",
value_name="estimate",
)
primary_ml_plot["effect_label"] = primary_ml_plot["effect_type"].map(
{
"gcomp_total_effect": "Total",
"natural_direct_effect": "Direct",
"natural_indirect_effect": "Indirect",
}
)
primary_ml_plot["model_label"] = primary_ml_plot["model_family"].str.upper()
fig, ax = plt.subplots(figsize=(11, 6))
sns.barplot(data=primary_ml_plot, x="estimate", y="model_label", hue="effect_label", ax=ax)
ax.axvline(0, color="black", linewidth=1)
ax.set_title("Cross-Fitted ML Mediation Effect Comparison")
ax.set_xlabel("Effect on future 7-day interactions")
ax.set_ylabel("Model family")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "21_crossfit_ml_effect_comparison.png", dpi=160, bbox_inches="tight")
plt.show()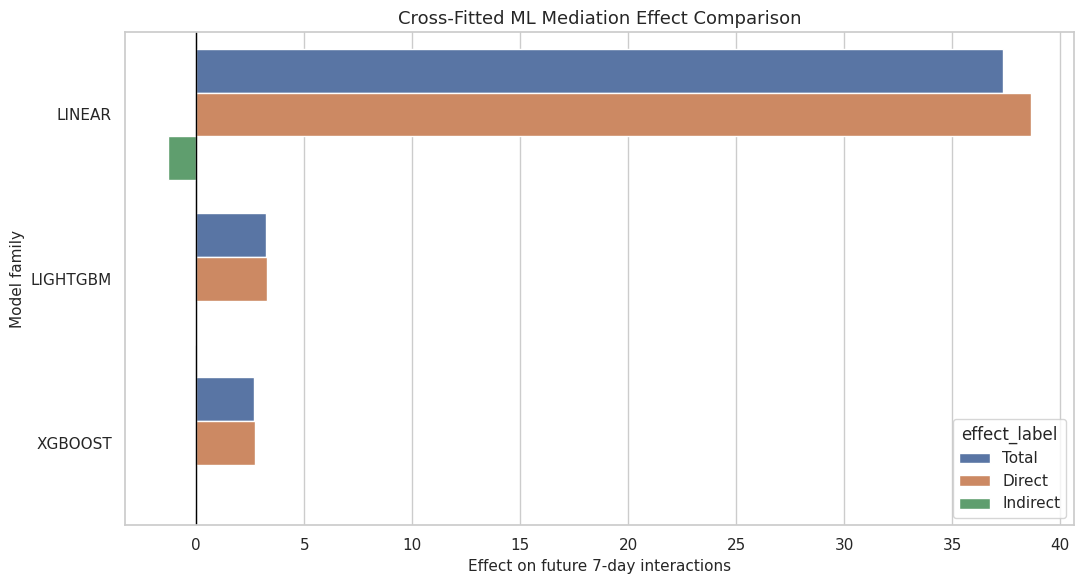
This comparison is the main advanced-model result. Agreement across Linear, LightGBM, and XGBoost is more reassuring than any one model alone.
12. Compare Predictive Performance
This cell compares out-of-fold predictive performance for the mediator and outcome nuisance models. Stronger prediction does not automatically mean a better causal estimate, but very weak prediction can make counterfactual estimates unstable.
performance_plot = ml_performance.copy()
performance_plot["model_label"] = performance_plot["model_family"].str.upper()
performance_plot["target_label"] = performance_plot["target"].str.title()
performance_plot["outcome_label"] = performance_plot["outcome"].map(
{PRIMARY_OUTCOME: "Future interactions", SECONDARY_OUTCOME: "Future play hours"}
)
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
for ax, metric in zip(axes, ["rmse", "r2"]):
sns.barplot(
data=performance_plot.query("outcome == @PRIMARY_OUTCOME"),
x=metric,
y="model_label",
hue="target_label",
ax=ax,
)
ax.set_title(f"Primary Outcome Nuisance {metric.upper()}")
ax.set_xlabel(metric.upper())
ax.set_ylabel("Model family")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "22_crossfit_nuisance_performance.png", dpi=160, bbox_inches="tight")
plt.show()
display(ml_performance.round(4))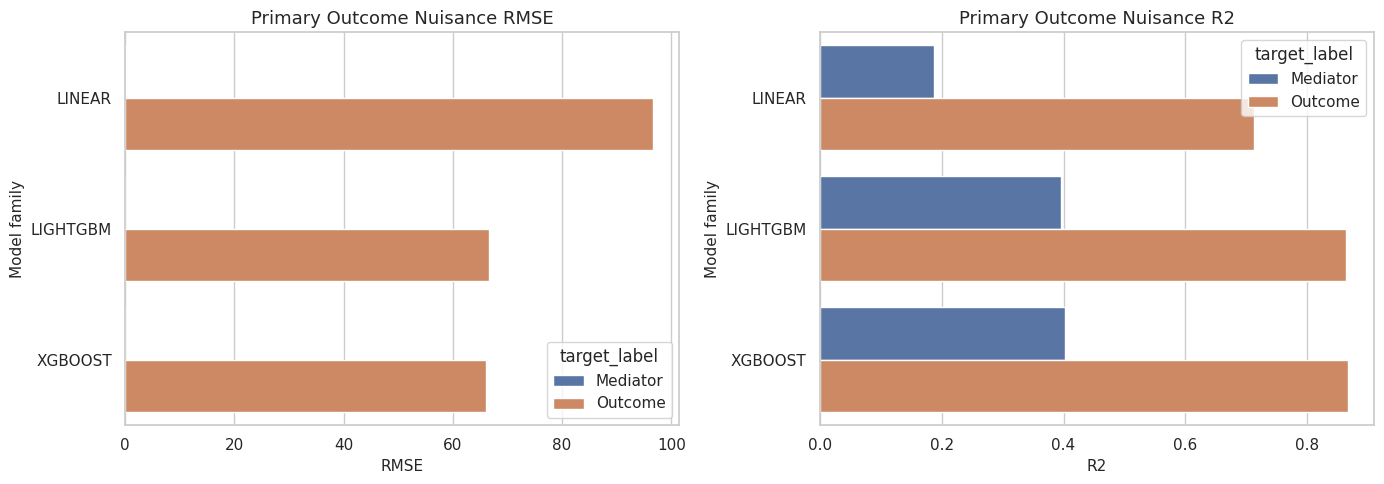
| model_family | outcome | target | rmse | mae | r2 | |
|---|---|---|---|---|---|---|
| 0 | linear | Y_future_interactions | mediator | 0.1015 | 0.0703 | 0.1878 |
| 1 | linear | Y_future_interactions | outcome | 96.6374 | 71.1050 | 0.7129 |
| 2 | lightgbm | Y_future_interactions | mediator | 0.0875 | 0.0629 | 0.3959 |
| 3 | lightgbm | Y_future_interactions | outcome | 66.7067 | 47.0941 | 0.8632 |
| 4 | xgboost | Y_future_interactions | mediator | 0.0870 | 0.0625 | 0.4026 |
| 5 | xgboost | Y_future_interactions | outcome | 66.0357 | 46.7289 | 0.8659 |
| 6 | linear | Y_future_play_hours | mediator | 0.1015 | 0.0703 | 0.1878 |
| 7 | linear | Y_future_play_hours | outcome | 0.3609 | 0.2428 | 0.4056 |
| 8 | lightgbm | Y_future_play_hours | mediator | 0.0875 | 0.0629 | 0.3959 |
| 9 | lightgbm | Y_future_play_hours | outcome | 0.2388 | 0.1672 | 0.7398 |
| 10 | xgboost | Y_future_play_hours | mediator | 0.0870 | 0.0625 | 0.4026 |
| 11 | xgboost | Y_future_play_hours | outcome | 0.2383 | 0.1694 | 0.7409 |
The performance table helps explain model behavior. If a flexible model changes the effect estimate but does not improve held-out performance, that change deserves caution.
13. Heterogeneity from Cross-Fitted Effects
This cell uses LightGBM’s row-level cross-fitted effects to summarize heterogeneity. The groups are based on pre-treatment information: prior activity, prior satisfaction, prior discovery, and user activity degree.
heterogeneity_base = analysis_panel[[
"user_id",
"event_date",
"prior_3day_interactions",
"prior_3day_high_satisfaction_share",
"prior_3day_discovery_candidate_share",
"user_active_degree",
]].copy()
heterogeneity_base["prior_activity_group"] = pd.qcut(
heterogeneity_base["prior_3day_interactions"].rank(method="first"),
q=3,
labels=["low prior activity", "medium prior activity", "high prior activity"],
)
heterogeneity_base["prior_satisfaction_group"] = pd.qcut(
heterogeneity_base["prior_3day_high_satisfaction_share"].rank(method="first"),
q=3,
labels=["low prior satisfaction", "medium prior satisfaction", "high prior satisfaction"],
)
heterogeneity_base["prior_discovery_group"] = pd.qcut(
heterogeneity_base["prior_3day_discovery_candidate_share"].rank(method="first"),
q=3,
labels=["low prior discovery", "medium prior discovery", "high prior discovery"],
)
lightgbm_row_effects = row_level_effects.query(
"model_family == 'lightgbm' and outcome == @PRIMARY_OUTCOME"
).merge(
heterogeneity_base,
on=["user_id", "event_date"],
how="left",
)
heterogeneity_rows = []
for segment_col in ["prior_activity_group", "prior_satisfaction_group", "prior_discovery_group", "user_active_degree"]:
summary = (
lightgbm_row_effects.groupby(segment_col, observed=True)
.agg(
user_days=("user_id", "size"),
users=("user_id", "nunique"),
total_effect=("ite_total", "mean"),
direct_effect=("ite_direct", "mean"),
indirect_effect=("ite_indirect", "mean"),
mediator_shift=("m1_hat", "mean"),
)
.reset_index()
.rename(columns={segment_col: "segment_value"})
)
summary["segment"] = segment_col
heterogeneity_rows.append(summary)
heterogeneity_effects = pd.concat(heterogeneity_rows, ignore_index=True)
# Convert mediator_shift from average M1 to the segment-level treatment-induced shift.
for segment_col in ["prior_activity_group", "prior_satisfaction_group", "prior_discovery_group", "user_active_degree"]:
mask = heterogeneity_effects["segment"].eq(segment_col)
segment_shift = (
lightgbm_row_effects.groupby(segment_col, observed=True)
.apply(lambda g: (g["m1_hat"] - g["m0_hat"]).mean(), include_groups=False)
.reset_index(name="true_mediator_shift")
.rename(columns={segment_col: "segment_value"})
)
heterogeneity_effects.loc[mask, "mediator_shift"] = heterogeneity_effects.loc[mask].merge(
segment_shift, on="segment_value", how="left"
)["true_mediator_shift"].to_numpy()
display(heterogeneity_effects.round(4))| segment_value | user_days | users | total_effect | direct_effect | indirect_effect | mediator_shift | segment | |
|---|---|---|---|---|---|---|---|---|
| 0 | low prior activity | 2733 | 133 | 2.1266 | 2.3286 | -0.2021 | 0.0195 | prior_activity_group |
| 1 | medium prior activity | 2733 | 133 | 3.9690 | 4.0530 | -0.0841 | 0.0177 | prior_activity_group |
| 2 | high prior activity | 2733 | 133 | 3.5253 | 3.4140 | 0.1112 | 0.0180 | prior_activity_group |
| 3 | low prior satisfaction | 2733 | 133 | 2.9903 | 2.8696 | 0.1208 | 0.0165 | prior_satisfaction_group |
| 4 | medium prior satisfaction | 2733 | 130 | 3.5907 | 3.7333 | -0.1426 | 0.0158 | prior_satisfaction_group |
| 5 | high prior satisfaction | 2733 | 113 | 3.0398 | 3.1928 | -0.1530 | 0.0229 | prior_satisfaction_group |
| 6 | low prior discovery | 2733 | 133 | 2.0592 | 2.1656 | -0.1064 | 0.0184 | prior_discovery_group |
| 7 | medium prior discovery | 2733 | 133 | 3.3925 | 3.3729 | 0.0196 | 0.0166 | prior_discovery_group |
| 8 | high prior discovery | 2733 | 133 | 4.1691 | 4.2572 | -0.0881 | 0.0201 | prior_discovery_group |
| 9 | UNKNOWN | 184 | 3 | 4.4181 | 4.5911 | -0.1731 | 0.0241 | user_active_degree |
| 10 | full_active | 7133 | 115 | 3.1915 | 3.2485 | -0.0570 | 0.0181 | user_active_degree |
| 11 | high_active | 882 | 15 | 3.0788 | 3.1241 | -0.0453 | 0.0195 | user_active_degree |
The heterogeneity table asks where the discovery effect is largest. Because these are subgroup summaries of cross-fitted predictions, they are descriptive signals for product targeting rather than separate randomized subgroup effects.
14. Plot Heterogeneous Effects
This cell plots LightGBM total effects by segment. It focuses on total effects because those are the most stable quantities in the earlier robustness checks.
heterogeneity_plot = heterogeneity_effects.copy()
heterogeneity_plot["segment_label"] = heterogeneity_plot["segment"].str.replace("_", " ").str.title()
heterogeneity_plot["segment_value"] = heterogeneity_plot["segment_value"].astype(str)
fig, axes = plt.subplots(2, 2, figsize=(15, 10))
axes = axes.flatten()
for ax, segment in zip(axes, heterogeneity_plot["segment"].unique()):
current = heterogeneity_plot.query("segment == @segment").copy()
sns.barplot(data=current, x="total_effect", y="segment_value", color="steelblue", ax=ax)
ax.axvline(0, color="black", linewidth=1)
ax.set_title(segment.replace("_", " ").title())
ax.set_xlabel("LightGBM total effect on future interactions")
ax.set_ylabel("")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "23_lightgbm_heterogeneous_effects.png", dpi=160, bbox_inches="tight")
plt.show()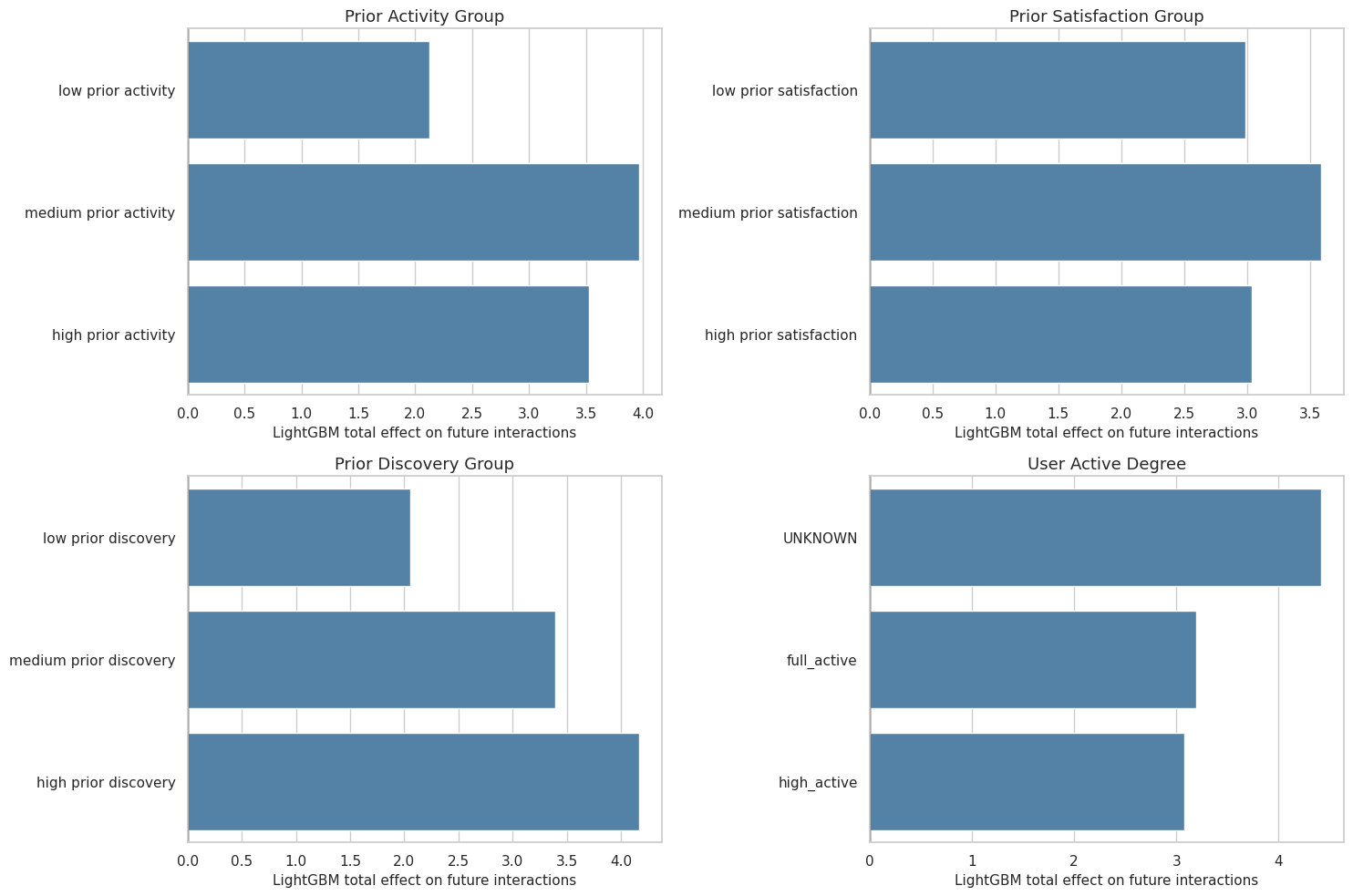
The heterogeneity plot makes the advanced notebook more product-relevant. It shows where the model expects discovery exposure to matter more or less.
15. Train Full Tree Models for Feature Importance
This cell trains full-sample LightGBM and XGBoost nuisance models for feature importance only. These models are not used for effect estimates; the effect estimates above are cross-fitted. The goal here is model auditability.
feature_importance_rows = []
full_design_m = treatment_design(X_covariates, A)
full_design_y = outcome_design(X_covariates, A, M)
importance_model_specs = [
("lightgbm", LGBMRegressor(
n_estimators=120,
learning_rate=0.05,
num_leaves=24,
subsample=0.85,
colsample_bytree=0.85,
random_state=RANDOM_SEED,
n_jobs=1,
verbose=-1,
)),
("xgboost", XGBRegressor(
n_estimators=120,
max_depth=3,
learning_rate=0.05,
subsample=0.85,
colsample_bytree=0.85,
objective="reg:squarederror",
tree_method="hist",
random_state=RANDOM_SEED,
n_jobs=1,
verbosity=0,
)),
]
for model_family, model in importance_model_specs:
mediator_model = clone(model)
outcome_model = clone(model)
fit_with_optional_weight(mediator_model, full_design_m, M, sample_weight=weights)
fit_with_optional_weight(outcome_model, full_design_y, Y, sample_weight=weights)
for target_name, fitted_model, design in [
("mediator", mediator_model, full_design_m),
("outcome", outcome_model, full_design_y),
]:
importances = getattr(fitted_model, "feature_importances_", None)
if importances is None:
continue
current = pd.DataFrame(
{
"model_family": model_family,
"target": target_name,
"feature": design.columns,
"importance": importances,
}
)
current["importance_share"] = current["importance"] / current["importance"].sum()
feature_importance_rows.append(current)
feature_importance = pd.concat(feature_importance_rows, ignore_index=True)
top_feature_importance = (
feature_importance.sort_values(["model_family", "target", "importance"], ascending=[True, True, False])
.groupby(["model_family", "target"], group_keys=False)
.head(12)
.reset_index(drop=True)
)
display(top_feature_importance.round(4))| model_family | target | feature | importance | importance_share | |
|---|---|---|---|---|---|
| 0 | lightgbm | mediator | calendar_day_index | 297.0000 | 0.1076 |
| 1 | lightgbm | mediator | prior_3day_high_satisfaction_share | 260.0000 | 0.0942 |
| 2 | lightgbm | mediator | lag_1_high_satisfaction_share | 249.0000 | 0.0902 |
| 3 | lightgbm | mediator | prior_3day_discovery_candidate_share | 187.0000 | 0.0678 |
| 4 | lightgbm | mediator | onehot_feat8 | 147.0000 | 0.0533 |
| 5 | lightgbm | mediator | lag_1_valid_play_share | 137.0000 | 0.0496 |
| 6 | lightgbm | mediator | prior_3day_valid_play_share | 122.0000 | 0.0442 |
| 7 | lightgbm | mediator | onehot_feat3 | 118.0000 | 0.0428 |
| 8 | lightgbm | mediator | register_days | 111.0000 | 0.0402 |
| 9 | lightgbm | mediator | prior_3day_total_play_duration_sec | 108.0000 | 0.0391 |
| 10 | lightgbm | mediator | lag_1_total_play_duration_sec | 103.0000 | 0.0373 |
| 11 | lightgbm | mediator | A_high_discovery | 101.0000 | 0.0366 |
| 12 | lightgbm | outcome | calendar_day_index | 592.0000 | 0.2145 |
| 13 | lightgbm | outcome | onehot_feat8 | 187.0000 | 0.0678 |
| 14 | lightgbm | outcome | prior_3day_discovery_candidate_share | 180.0000 | 0.0652 |
| 15 | lightgbm | outcome | prior_3day_interactions | 157.0000 | 0.0569 |
| 16 | lightgbm | outcome | register_days | 148.0000 | 0.0536 |
| 17 | lightgbm | outcome | onehot_feat7 | 126.0000 | 0.0457 |
| 18 | lightgbm | outcome | prior_3day_high_satisfaction_share | 121.0000 | 0.0438 |
| 19 | lightgbm | outcome | onehot_feat3 | 120.0000 | 0.0435 |
| 20 | lightgbm | outcome | follow_user_num | 108.0000 | 0.0391 |
| 21 | lightgbm | outcome | prior_3day_total_play_duration_sec | 105.0000 | 0.0380 |
| 22 | lightgbm | outcome | prior_3day_valid_play_share | 103.0000 | 0.0373 |
| 23 | lightgbm | outcome | fans_user_num | 87.0000 | 0.0315 |
| 24 | xgboost | mediator | prior_3day_high_satisfaction_share | 0.1697 | 0.1697 |
| 25 | xgboost | mediator | lag_1_high_satisfaction_share | 0.0686 | 0.0686 |
| 26 | xgboost | mediator | lag_1_valid_play_share | 0.0570 | 0.0570 |
| 27 | xgboost | mediator | prior_3day_valid_play_share | 0.0436 | 0.0436 |
| 28 | xgboost | mediator | lag_1_interactions | 0.0432 | 0.0432 |
| 29 | xgboost | mediator | onehot_feat11 | 0.0416 | 0.0416 |
| 30 | xgboost | mediator | fans_user_num_range_1_10 | 0.0381 | 0.0381 |
| 31 | xgboost | mediator | user_active_degree_high_active | 0.0330 | 0.0330 |
| 32 | xgboost | mediator | prior_3day_active_day | 0.0312 | 0.0312 |
| 33 | xgboost | mediator | onehot_feat6 | 0.0262 | 0.0262 |
| 34 | xgboost | mediator | lag_1_total_play_duration_sec | 0.0261 | 0.0261 |
| 35 | xgboost | mediator | A_high_discovery | 0.0227 | 0.0227 |
| 36 | xgboost | outcome | calendar_day_index | 0.3093 | 0.3093 |
| 37 | xgboost | outcome | prior_3day_active_day | 0.1905 | 0.1905 |
| 38 | xgboost | outcome | prior_3day_discovery_candidate_share | 0.1517 | 0.1517 |
| 39 | xgboost | outcome | prior_3day_interactions | 0.0373 | 0.0373 |
| 40 | xgboost | outcome | lag_1_interactions | 0.0202 | 0.0202 |
| 41 | xgboost | outcome | lag_1_discovery_candidate_share | 0.0190 | 0.0190 |
| 42 | xgboost | outcome | recent_activity_score | 0.0169 | 0.0169 |
| 43 | xgboost | outcome | user_active_degree_high_active | 0.0161 | 0.0161 |
| 44 | xgboost | outcome | prior_3day_total_play_duration_sec | 0.0145 | 0.0145 |
| 45 | xgboost | outcome | register_days_range_61_90 | 0.0141 | 0.0141 |
| 46 | xgboost | outcome | fans_user_num | 0.0136 | 0.0136 |
| 47 | xgboost | outcome | onehot_feat14 | 0.0117 | 0.0117 |
The feature-importance audit helps reveal whether models are mostly using treatment, mediator, calendar time, prior activity, or profile signals. That matters because an advanced model can look impressive while leaning heavily on confounding proxies.
16. Plot Feature Importance
This cell plots the top features for the LightGBM nuisance models. LightGBM is used here because its importances are usually stable and easy to read.
importance_plot = top_feature_importance.query("model_family == 'lightgbm'").copy()
importance_plot["feature_label"] = importance_plot["feature"].str.replace("_", " ").str.slice(0, 45)
importance_plot["target_label"] = importance_plot["target"].str.title()
fig, axes = plt.subplots(1, 2, figsize=(16, 6))
for ax, target in zip(axes, ["mediator", "outcome"]):
current = importance_plot.query("target == @target").sort_values("importance", ascending=True)
sns.barplot(data=current, x="importance", y="feature_label", color="steelblue", ax=ax)
ax.set_title(f"LightGBM {target.title()} Model Features")
ax.set_xlabel("Feature importance")
ax.set_ylabel("")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "24_lightgbm_feature_importance.png", dpi=160, bbox_inches="tight")
plt.show()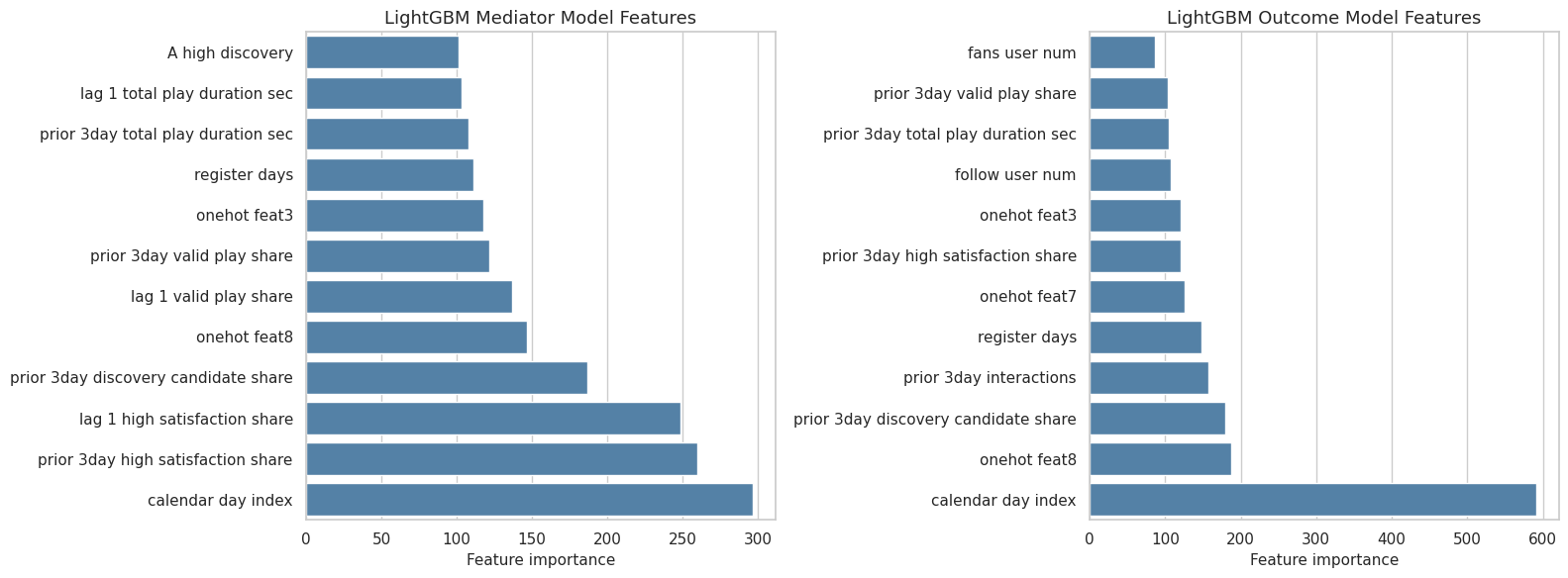
The feature plot is an audit layer, not an explanation of causality. It tells us what the prediction functions used, which is useful for debugging and for communicating model behavior.
17. Advanced Model Summary
This cell combines the linear g-computation result from notebook 04, SEM-style paths, and cross-fitted ML estimates into one comparison table. The table is the main handoff to the final report.
linear_reference = effect_summary.query(
"outcome == @PRIMARY_OUTCOME and estimand in ['gcomp_total_effect', 'natural_direct_effect', 'natural_indirect_effect']"
).pivot_table(index="outcome", columns="estimand", values="estimate").reset_index()
linear_reference["model_family"] = "linear_reference_from_notebook_04"
linear_reference = linear_reference.rename(
columns={
"gcomp_total_effect": "total_effect",
"natural_direct_effect": "direct_effect",
"natural_indirect_effect": "indirect_effect",
}
)[["model_family", "total_effect", "direct_effect", "indirect_effect"]]
sem_reference = pd.DataFrame(
[
{
"model_family": "sem_style_path_model",
"total_effect": sem_path_results.loc[0, "total_path_c_prime_plus_indirect"],
"direct_effect": sem_path_results.loc[0, "c_prime_A_to_Y"],
"indirect_effect": sem_path_results.loc[0, "indirect_alpha_beta"],
}
]
)
ml_reference = ml_effects.query("outcome == @PRIMARY_OUTCOME").rename(
columns={
"gcomp_total_effect": "total_effect",
"natural_direct_effect": "direct_effect",
"natural_indirect_effect": "indirect_effect",
}
)[["model_family", "total_effect", "direct_effect", "indirect_effect", "mediator_shift", "relative_total_effect"]]
advanced_model_summary = pd.concat(
[linear_reference, sem_reference, ml_reference],
ignore_index=True,
sort=False,
)
advanced_model_summary["indirect_to_total_ratio"] = (
advanced_model_summary["indirect_effect"] / advanced_model_summary["total_effect"].replace(0, np.nan)
)
advanced_model_summary["total_effect_positive"] = advanced_model_summary["total_effect"] > 0
advanced_model_summary["indirect_effect_small_abs_lt_5"] = advanced_model_summary["indirect_effect"].abs() < 5
display(advanced_model_summary.round(4))| model_family | total_effect | direct_effect | indirect_effect | mediator_shift | relative_total_effect | indirect_to_total_ratio | total_effect_positive | indirect_effect_small_abs_lt_5 | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | linear_reference_from_notebook_04 | 37.2781 | 38.5858 | -1.3077 | NaN | NaN | -0.0351 | True | True |
| 1 | sem_style_path_model | 37.3139 | 38.4311 | -1.1172 | NaN | NaN | -0.0299 | True | True |
| 2 | linear | 37.3421 | 38.6464 | -1.3043 | 0.0231 | 0.1156 | -0.0349 | True | True |
| 3 | lightgbm | 3.2069 | 3.2652 | -0.0583 | 0.0184 | 0.0095 | -0.0182 | True | True |
| 4 | xgboost | 2.6829 | 2.7279 | -0.0450 | 0.0138 | 0.0079 | -0.0168 | True | True |
The comparison table answers the advanced-model question directly. If the total effect is positive across model families and the indirect effect remains small, the notebook 04 result is strengthened.
18. Plot Advanced Model Summary
This cell plots total, direct, and indirect effects across the reference, SEM-style, and cross-fitted ML approaches.
summary_plot = advanced_model_summary.melt(
id_vars=["model_family"],
value_vars=["total_effect", "direct_effect", "indirect_effect"],
var_name="effect_type",
value_name="estimate",
)
summary_plot["effect_label"] = summary_plot["effect_type"].map(
{
"total_effect": "Total",
"direct_effect": "Direct",
"indirect_effect": "Indirect",
}
)
summary_plot["model_label"] = summary_plot["model_family"].str.replace("_", " ").str.title()
fig, ax = plt.subplots(figsize=(13, 7))
sns.barplot(data=summary_plot, x="estimate", y="model_label", hue="effect_label", ax=ax)
ax.axvline(0, color="black", linewidth=1)
ax.set_title("Advanced Model Comparison for Discovery Mediation")
ax.set_xlabel("Effect on future 7-day interactions")
ax.set_ylabel("Model approach")
plt.tight_layout()
fig.savefig(FIGURE_DIR / "25_advanced_model_summary.png", dpi=160, bbox_inches="tight")
plt.show()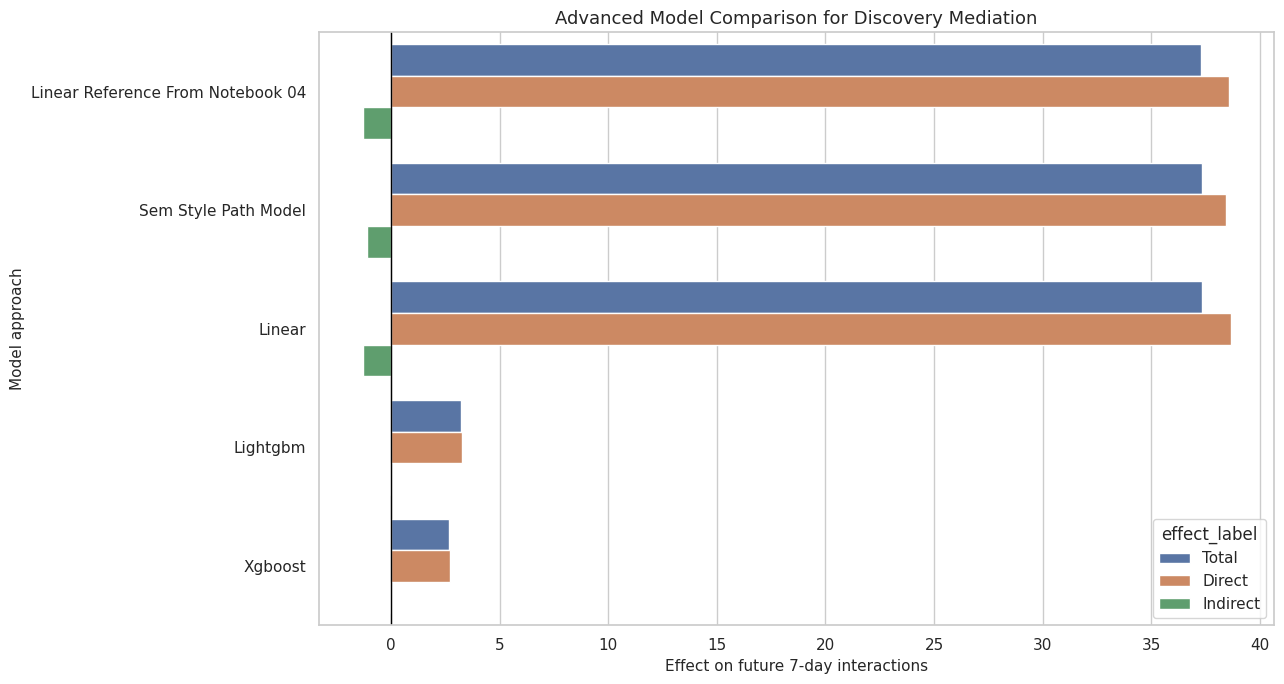
This final figure is the high-level advanced-model takeaway. It lets a reader see whether the main story depends on the modeling approach.
19. Save Advanced Model Artifacts
This cell saves the SEM path results, bootstrap path samples, ML effect estimates, model performance, heterogeneity table, feature importance table, and advanced summary table.
sem_path_summary.to_csv(SEM_PATH_OUTPUT, index=False)
sem_bootstrap.to_csv(SEM_BOOTSTRAP_OUTPUT, index=False)
ml_effects.to_csv(ML_EFFECTS_OUTPUT, index=False)
ml_performance.to_csv(ML_PERFORMANCE_OUTPUT, index=False)
heterogeneity_effects.to_csv(HETEROGENEITY_OUTPUT, index=False)
feature_importance.to_csv(FEATURE_IMPORTANCE_OUTPUT, index=False)
advanced_model_summary.to_csv(ADVANCED_SUMMARY_OUTPUT, index=False)
sem_path_summary.to_csv(TABLE_DIR / "advanced_sem_path_results.csv", index=False)
ml_effects.to_csv(TABLE_DIR / "advanced_ml_effects.csv", index=False)
ml_performance.to_csv(TABLE_DIR / "advanced_ml_performance.csv", index=False)
heterogeneity_effects.to_csv(TABLE_DIR / "advanced_heterogeneity.csv", index=False)
top_feature_importance.to_csv(TABLE_DIR / "advanced_feature_importance_top.csv", index=False)
advanced_model_summary.to_csv(TABLE_DIR / "advanced_model_summary.csv", index=False)
saved_outputs = pd.DataFrame(
{
"artifact": [
"sem_path_results",
"sem_bootstrap",
"ml_effects",
"ml_performance",
"heterogeneity_effects",
"feature_importance",
"advanced_model_summary",
],
"path": [
str(SEM_PATH_OUTPUT),
str(SEM_BOOTSTRAP_OUTPUT),
str(ML_EFFECTS_OUTPUT),
str(ML_PERFORMANCE_OUTPUT),
str(HETEROGENEITY_OUTPUT),
str(FEATURE_IMPORTANCE_OUTPUT),
str(ADVANCED_SUMMARY_OUTPUT),
],
}
)
display(saved_outputs)| artifact | path | |
|---|---|---|
| 0 | sem_path_results | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_sem_path_results.csv |
| 1 | sem_bootstrap | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_sem_bootstrap.csv |
| 2 | ml_effects | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_advanced_ml_effects.csv |
| 3 | ml_performance | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_advanced_ml_performance.csv |
| 4 | heterogeneity_effects | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_advanced_heterogeneity.csv |
| 5 | feature_importance | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_advanced_feature_importance.csv |
| 6 | advanced_model_summary | /home/apex/Documents/ranking_sys/data/processed/kuairec_discovery_quality_advanced_model_summary.csv |
The saved tables give the final report enough material to compare simple and advanced estimators without rerunning the models.
20. Notebook Takeaways
This notebook added advanced model checks to the discovery-quality mediation workflow:
- SEM-style path modeling gives a compact path-coefficient version of the mediation story.
- Cross-fitted Linear, LightGBM, and XGBoost nuisance models test whether the decomposition survives nonlinear prediction functions.
- Heterogeneity summaries show where discovery exposure appears more or less valuable.
- Feature-importance tables audit which variables the tree-based nuisance models rely on.
- The final comparison table tells the final report whether the main result is stable across modeling approaches.
The next notebook should be the final report and figures notebook, using the simple, robustness, and advanced-model outputs together.