import importlib.util
import json
import sys
import textwrap
import warnings
from pathlib import Path
from typing import Literal
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import statsmodels.formula.api as smf
import torch
from IPython.display import Markdown, display
from pydantic import BaseModel, Field
from sklearn.linear_model import LinearRegression
warnings.filterwarnings('ignore', category=FutureWarning)16. AI for Sensitivity Analysis
Sensitivity analysis asks a disciplined question:
How wrong would our causal conclusion be under plausible violations of the assumptions?
This is different from asking whether a coefficient is statistically significant. An observational estimate can be precisely estimated and still be badly biased. A sensitivity analysis makes design fragility visible: unobserved confounding, omitted variables, weak overlap, measurement error, placebo failures, and bad-control risks.
This notebook shows how AI can help structure sensitivity analysis without replacing it. Python computes the evidence. The LLM helps organize interpretation and communication. The audit layer checks whether the model overclaims robustness.
Learning Goals
By the end of this notebook you should be able to:
- Explain why sensitivity analysis is part of the causal result, not an optional appendix.
- Estimate a baseline observational effect and compare it with an oracle estimate in synthetic data.
- Use observed covariates as benchmarks for omitted-variable sensitivity.
- Build a residual-confounding sensitivity grid that asks how strong an unobserved confounder would need to be.
- Use placebo outcomes to detect design fragility.
- Assemble a machine-readable sensitivity report.
- Use local LLMs to summarize sensitivity evidence while auditing them for overconfident robustness claims.
Live Model Note
This course treats LLM behavior as an empirical object. These notebooks may include live local-model calls, so outputs can vary across model versions, hardware, decoding settings, prompt wording, package versions, and reruns. That instability is part of the lesson: AI-assisted causal work needs deterministic checks, structured outputs, model comparison, repair logic, and human review.
Sensitivity-analysis notebooks are brittle because the interpretation is subtle. A model may correctly list sensitivity checks but still say “robust” too quickly. It may ignore a placebo failure, confuse oracle synthetic columns with real evidence, or treat a benchmark calculation as proof. We will make those risks explicit and score the model’s interpretation against deterministic gates.
1. Setup
The deterministic part of this notebook uses a synthetic observational example with a known hidden confounder. In real data we would not observe the hidden confounder; here we keep it for teaching and oracle comparisons.
RUN_LIVE_LOCAL_LLM = True
RUN_FULL_MODEL_COMPARISON = True
RUN_SCHEMA_REPAIR_RETRY = True
LOCAL_SMOKE_TEST_MODEL = 'Qwen/Qwen2.5-0.5B-Instruct'
LOCAL_FAST_MODEL = 'Qwen/Qwen2.5-7B-Instruct'
LOCAL_STRONG_MODEL = 'Qwen/Qwen2.5-14B-Instruct'
LOCAL_SCALE_MODEL = 'Qwen/Qwen2.5-32B-Instruct'
LOCAL_ALT_REASONING_MODEL = 'microsoft/Phi-3.5-mini-instruct'
LOCAL_ALT_OPEN_MODEL = 'mistralai/Mistral-7B-Instruct-v0.3'
LOCAL_MISTRAL_SMALL_MODEL = 'mistralai/Mistral-Small-3.1-24B-Instruct-2503'
LOCAL_GEMMA_MODEL = 'google/gemma-3-27b-it'
LOCAL_LLAMA_MODEL = 'meta-llama/Meta-Llama-3.1-8B-Instruct'
MODEL_ID = LOCAL_STRONG_MODEL
MAX_NEW_TOKENS = 1900
TEMPERATURE = 0.0
SEED = 216
MODEL_COMPARISON_CASE_LIMIT = 2
MODELS_TO_COMPARE = [
('Qwen 0.5B', LOCAL_SMOKE_TEST_MODEL, 'pipeline smoke test'),
('Qwen 7B', LOCAL_FAST_MODEL, 'fast default'),
('Qwen 14B', LOCAL_STRONG_MODEL, 'strong local analysis'),
('Qwen 32B', LOCAL_SCALE_MODEL, 'scale comparison'),
('Phi mini', LOCAL_ALT_REASONING_MODEL, 'compact non-Qwen comparison'),
('Mistral 7B', LOCAL_ALT_OPEN_MODEL, '7B model-family comparison'),
('Mistral Small 24B', LOCAL_MISTRAL_SMALL_MODEL, 'strong non-Qwen comparison'),
('Gemma 3 27B', LOCAL_GEMMA_MODEL, 'large non-Qwen comparison'),
('Llama 3.1 8B', LOCAL_LLAMA_MODEL, 'industry-standard instruct baseline'),
]
np.random.seed(SEED)
sns.set_theme(style='whitegrid', context='notebook')
pd.set_option('display.max_colwidth', 160)def has_package(module_name):
return importlib.util.find_spec(module_name) is not None
package_status = pd.DataFrame(
[
{'package': 'pandas', 'available': has_package('pandas'), 'used_for': 'sensitivity tables'},
{'package': 'statsmodels', 'available': has_package('statsmodels'), 'used_for': 'effect and placebo regressions'},
{'package': 'sklearn', 'available': has_package('sklearn'), 'used_for': 'residualization helpers'},
{'package': 'seaborn', 'available': has_package('seaborn'), 'used_for': 'sensitivity plots'},
{'package': 'pydantic', 'available': has_package('pydantic'), 'used_for': 'structured AI sensitivity review'},
{'package': 'transformers', 'available': has_package('transformers'), 'used_for': 'optional local LLM review'},
{'package': 'torch', 'available': has_package('torch'), 'used_for': 'GPU inference if live LLMs are enabled'},
]
)
print(f'CUDA available to this kernel: {torch.cuda.is_available()}')
package_statusCUDA available to this kernel: True| package | available | used_for | |
|---|---|---|---|
| 0 | pandas | True | sensitivity tables |
| 1 | statsmodels | True | effect and placebo regressions |
| 2 | sklearn | True | residualization helpers |
| 3 | seaborn | True | sensitivity plots |
| 4 | pydantic | True | structured AI sensitivity review |
| 5 | transformers | True | optional local LLM review |
| 6 | torch | True | GPU inference if live LLMs are enabled |
2. What Sensitivity Analysis Is For
Sensitivity analysis is not pessimism. It is disciplined honesty.
A standard adjusted estimate says:
Under the stated assumptions, this is the estimated effect.
A sensitivity analysis adds:
If the assumptions are wrong in these specific ways, this is how the estimate would move.
The second sentence is often the difference between a defensible causal report and a brittle dashboard claim.
sensitivity_components = pd.DataFrame(
[
{
'component': 'Benchmark confounders',
'question': 'How much does the treatment effect move when important observed covariates are omitted?',
'what it teaches': 'Plausible scale for omitted-variable bias.',
},
{
'component': 'Residual confounding grid',
'question': 'How strong would a hidden confounder need to be with residual treatment and outcome?',
'what it teaches': 'Tipping points for sign reversal or loss of practical significance.',
},
{
'component': 'Placebo outcome',
'question': 'Does treatment appear to affect an outcome measured before treatment?',
'what it teaches': 'Evidence of residual confounding or selection.',
},
{
'component': 'Oracle comparison in simulation',
'question': 'How far is feasible adjustment from the estimate that includes hidden truth variables?',
'what it teaches': 'What the observed analysis is missing in a controlled lab.',
},
{
'component': 'Communication warning',
'question': 'What should stakeholders not conclude from this estimate?',
'what it teaches': 'Prevents robustness language from becoming overclaiming.',
},
]
)
sensitivity_components| component | question | what it teaches | |
|---|---|---|---|
| 0 | Benchmark confounders | How much does the treatment effect move when important observed covariates are omitted? | Plausible scale for omitted-variable bias. |
| 1 | Residual confounding grid | How strong would a hidden confounder need to be with residual treatment and outcome? | Tipping points for sign reversal or loss of practical significance. |
| 2 | Placebo outcome | Does treatment appear to affect an outcome measured before treatment? | Evidence of residual confounding or selection. |
| 3 | Oracle comparison in simulation | How far is feasible adjustment from the estimate that includes hidden truth variables? | What the observed analysis is missing in a controlled lab. |
| 4 | Communication warning | What should stakeholders not conclude from this estimate? | Prevents robustness language from becoming overclaiming. |
Discussion
No single sensitivity method is universal. The right set depends on the design: unmeasured confounding for observational adjustment, pre-trends for DiD, exclusion violations for IV, manipulation for RDD, interference for experiments, and logging-policy overlap for off-policy evaluation.
3. Running Example: Onboarding Specialist Assignment
A SaaS company assigns some customers to a specialist onboarding program. The business asks:
What is the effect of specialist onboarding on 90-day net revenue?
Assignment is observational. Managers prioritize accounts with high value, high risk, and signs of executive attention. Some of that attention is recorded; some is not. We simulate the hidden variable so we can teach sensitivity analysis.
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def simulate_onboarding_data(n=5200, seed=SEED, hidden_strength=0.65, overlap_noise=0.75):
rng = np.random.default_rng(seed)
segment = rng.choice(['startup', 'midmarket', 'enterprise'], size=n, p=[0.46, 0.38, 0.16])
region = rng.choice(['Americas', 'EMEA', 'APAC'], size=n, p=[0.48, 0.33, 0.19])
tenure_months = rng.gamma(shape=2.5, scale=7.5, size=n).clip(1, 72)
baseline_usage = rng.normal(0, 1, size=n)
prior_support_tickets = rng.poisson(np.exp(-0.20 - 0.18 * baseline_usage + 0.24 * (segment == 'enterprise')), size=n)
contract_value = np.exp(3.3 + 0.45 * (segment == 'enterprise') + 0.18 * baseline_usage + rng.normal(0, 0.42, size=n))
observed_health = 0.62 * baseline_usage + 0.018 * tenure_months - 0.24 * prior_support_tickets + 0.18 * (region == 'Americas') + rng.normal(0, 0.58, size=n)
risk_score = -0.86 * observed_health + 0.30 * prior_support_tickets + 0.12 * np.log1p(contract_value) + rng.normal(0, 0.45, size=n)
executive_attention = rng.normal(0, 1, size=n)
prior_period_revenue = (
24
+ 0.34 * contract_value
+ 7.0 * observed_health
+ hidden_strength * 7.5 * executive_attention
+ rng.normal(0, 18, size=n)
)
assignment_logit = (
-0.25
+ 1.05 * risk_score
+ 0.24 * np.log1p(contract_value)
+ 0.18 * (segment == 'enterprise')
+ hidden_strength * 0.85 * executive_attention
+ rng.normal(0, overlap_noise, size=n)
)
onboarding_specialist = rng.binomial(1, sigmoid(assignment_logit))
true_effect = 7.0 + 2.0 * sigmoid(risk_score) + 1.2 * (segment == 'enterprise')
net_revenue_90d = (
30
+ 0.42 * contract_value
+ 10.0 * observed_health
- 2.0 * prior_support_tickets
+ hidden_strength * 9.0 * executive_attention
+ true_effect * onboarding_specialist
+ rng.normal(0, 22, size=n)
)
product_adoption_30d = 0.45 * baseline_usage + 0.55 * onboarding_specialist + 0.30 * executive_attention + rng.normal(0, 1, size=n)
expansion_pipeline_120d = net_revenue_90d + 0.20 * contract_value + 12 * onboarding_specialist + rng.normal(0, 20, size=n)
df = pd.DataFrame(
{
'account_id': [f'A{idx:05d}' for idx in range(n)],
'segment': segment,
'region': region,
'tenure_months': tenure_months,
'baseline_usage': baseline_usage,
'prior_support_tickets': prior_support_tickets,
'contract_value': contract_value,
'observed_health': observed_health,
'risk_score': risk_score,
'executive_attention': executive_attention,
'prior_period_revenue': prior_period_revenue,
'onboarding_specialist': onboarding_specialist,
'true_effect': true_effect,
'net_revenue_90d': net_revenue_90d,
'product_adoption_30d': product_adoption_30d,
'expansion_pipeline_120d': expansion_pipeline_120d,
}
)
truth = {
'true_ate': float(df['true_effect'].mean()),
'true_att': float(df.loc[df['onboarding_specialist'].eq(1), 'true_effect'].mean()),
'treatment_rate': float(df['onboarding_specialist'].mean()),
'hidden_strength': hidden_strength,
}
return df, truth
raw, truth = simulate_onboarding_data()
raw.head()| account_id | segment | region | tenure_months | baseline_usage | prior_support_tickets | contract_value | observed_health | risk_score | executive_attention | prior_period_revenue | onboarding_specialist | true_effect | net_revenue_90d | product_adoption_30d | expansion_pipeline_120d | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | A00000 | enterprise | Americas | 19.767521 | 1.440981 | 1 | 42.053951 | 0.372740 | 0.646366 | -0.881831 | 35.639973 | 0 | 9.512382 | 73.822233 | 1.491241 | 125.168395 |
| 1 | A00001 | midmarket | Americas | 2.714544 | -0.799302 | 0 | 37.342631 | 0.143256 | 0.532391 | 0.166796 | 24.695390 | 1 | 8.260081 | 96.890555 | -0.940938 | 108.780270 |
| 2 | A00002 | startup | EMEA | 54.283982 | -1.945033 | 1 | 10.483906 | 0.664454 | -0.882747 | -1.052783 | 1.203831 | 1 | 7.585218 | 38.893951 | -0.752022 | 43.501021 |
| 3 | A00003 | midmarket | Americas | 20.707722 | -0.813210 | 3 | 12.673929 | -0.047837 | 2.111590 | 0.039967 | 43.765324 | 1 | 8.784049 | 30.239183 | -1.263760 | 37.587590 |
| 4 | A00004 | midmarket | EMEA | 17.547371 | -1.420773 | 1 | 38.000833 | -0.279677 | 0.550063 | -1.428694 | 14.994348 | 0 | 8.268300 | 78.246247 | -0.659701 | 56.833462 |
truth{'true_ate': 8.388275446972925,
'true_att': 8.50067894836023,
'treatment_rate': 0.685,
'hidden_strength': 0.65}4. Analysis Contract and Variable Roles
The sensitivity report is tied to a specific estimand and variable role map. The hidden variable is included only because this is a synthetic teaching notebook.
analysis_contract = {
'analysis_id': 'onboarding_specialist_sensitivity_v1',
'estimand': 'ATE of onboarding_specialist on net_revenue_90d among eligible accounts',
'treatment': 'onboarding_specialist',
'outcome': 'net_revenue_90d',
'placebo_outcome': 'prior_period_revenue',
'allowed_covariates': [
'segment',
'region',
'tenure_months',
'baseline_usage',
'prior_support_tickets',
'contract_value',
'observed_health',
'risk_score',
],
'categorical_covariates': ['segment', 'region'],
'oracle_hidden_confounders': ['executive_attention'],
'forbidden_variables': ['product_adoption_30d', 'expansion_pipeline_120d', 'true_effect'],
'diagnostic_thresholds': {
'placebo_caution_revenue_units': 4.0,
'tipping_point_distance_caution': 0.35,
'benchmark_shift_caution_ratio': 1.0,
},
}
variable_roles = pd.DataFrame(
[{'variable': var, 'role': 'allowed pre-treatment covariate'} for var in analysis_contract['allowed_covariates']]
+ [{'variable': analysis_contract['treatment'], 'role': 'treatment'}]
+ [{'variable': analysis_contract['outcome'], 'role': 'outcome'}]
+ [{'variable': analysis_contract['placebo_outcome'], 'role': 'pre-treatment placebo outcome'}]
+ [{'variable': var, 'role': 'oracle hidden confounder for teaching only'} for var in analysis_contract['oracle_hidden_confounders']]
+ [{'variable': var, 'role': 'forbidden post-treatment/future/oracle variable'} for var in analysis_contract['forbidden_variables']]
)
variable_roles| variable | role | |
|---|---|---|
| 0 | segment | allowed pre-treatment covariate |
| 1 | region | allowed pre-treatment covariate |
| 2 | tenure_months | allowed pre-treatment covariate |
| 3 | baseline_usage | allowed pre-treatment covariate |
| 4 | prior_support_tickets | allowed pre-treatment covariate |
| 5 | contract_value | allowed pre-treatment covariate |
| 6 | observed_health | allowed pre-treatment covariate |
| 7 | risk_score | allowed pre-treatment covariate |
| 8 | onboarding_specialist | treatment |
| 9 | net_revenue_90d | outcome |
| 10 | prior_period_revenue | pre-treatment placebo outcome |
| 11 | executive_attention | oracle hidden confounder for teaching only |
| 12 | product_adoption_30d | forbidden post-treatment/future/oracle variable |
| 13 | expansion_pipeline_120d | forbidden post-treatment/future/oracle variable |
| 14 | true_effect | forbidden post-treatment/future/oracle variable |
Discussion
The forbidden variables are important. product_adoption_30d is post-treatment. expansion_pipeline_120d is future-looking. true_effect is simulation truth. Sensitivity analysis should not quietly turn these into controls.
5. Baseline, Oracle, and Placebo Estimates
We estimate three models:
- A feasible adjusted model using observed pre-treatment covariates.
- An oracle model that additionally controls for the hidden confounder. This is for teaching only.
- A placebo model using the same adjustment set but with a pre-treatment outcome.
def formula_term(variable, categorical):
return f'C({variable})' if variable in categorical else variable
def build_formula(outcome, treatment, covariates, categorical):
terms = [formula_term(var, categorical) for var in covariates]
return f"{outcome} ~ {treatment} + " + ' + '.join(terms)
def fit_effect_model(df, outcome, treatment, covariates, categorical):
formula = build_formula(outcome, treatment, covariates, categorical)
fitted = smf.ols(formula, data=df).fit(cov_type='HC3')
estimate = fitted.params[treatment]
se = fitted.bse[treatment]
return {
'formula': formula,
'estimate': float(estimate),
'robust_se': float(se),
'ci_low': float(estimate - 1.96 * se),
'ci_high': float(estimate + 1.96 * se),
'p_value': float(fitted.pvalues[treatment]),
'rows': int(fitted.nobs),
'model': fitted,
}
observed_result = fit_effect_model(
raw,
analysis_contract['outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'],
analysis_contract['categorical_covariates'],
)
oracle_result = fit_effect_model(
raw,
analysis_contract['outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'] + analysis_contract['oracle_hidden_confounders'],
analysis_contract['categorical_covariates'],
)
placebo_result = fit_effect_model(
raw,
analysis_contract['placebo_outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'],
analysis_contract['categorical_covariates'],
)
estimate_table = pd.DataFrame(
[
{'model': 'observed adjusted', 'target': 'outcome', **{k: v for k, v in observed_result.items() if k != 'model'}},
{'model': 'oracle hidden-confounder adjusted', 'target': 'outcome', **{k: v for k, v in oracle_result.items() if k != 'model'}},
{'model': 'observed adjusted placebo', 'target': 'placebo', **{k: v for k, v in placebo_result.items() if k != 'model'}},
]
)
estimate_table[['model', 'target', 'estimate', 'robust_se', 'ci_low', 'ci_high', 'p_value', 'rows']]| model | target | estimate | robust_se | ci_low | ci_high | p_value | rows | |
|---|---|---|---|---|---|---|---|---|
| 0 | observed adjusted | outcome | 9.757901 | 0.729626 | 8.327834 | 11.187967 | 8.597864e-41 | 5200 |
| 1 | oracle hidden-confounder adjusted | outcome | 7.336248 | 0.717084 | 5.930764 | 8.741732 | 1.445147e-24 | 5200 |
| 2 | observed adjusted placebo | placebo | 2.194321 | 0.604380 | 1.009736 | 3.378906 | 2.826560e-04 | 5200 |
fig, ax = plt.subplots(figsize=(9, 3.8))
plot_estimates = estimate_table.query("target == 'outcome'").copy()
ax.errorbar(
plot_estimates['estimate'],
plot_estimates['model'],
xerr=[plot_estimates['estimate'] - plot_estimates['ci_low'], plot_estimates['ci_high'] - plot_estimates['estimate']],
fmt='o',
color='#4477AA',
ecolor='#777777',
capsize=4,
)
ax.axvline(truth['true_ate'], color='#CC3311', linestyle='--', linewidth=2, label='true ATE')
ax.axvline(0, color='black', linewidth=1)
ax.set_title('Observed versus oracle sensitivity check')
ax.set_xlabel('Estimated effect on 90-day net revenue')
ax.set_ylabel('')
ax.legend()
plt.tight_layout()
plt.show()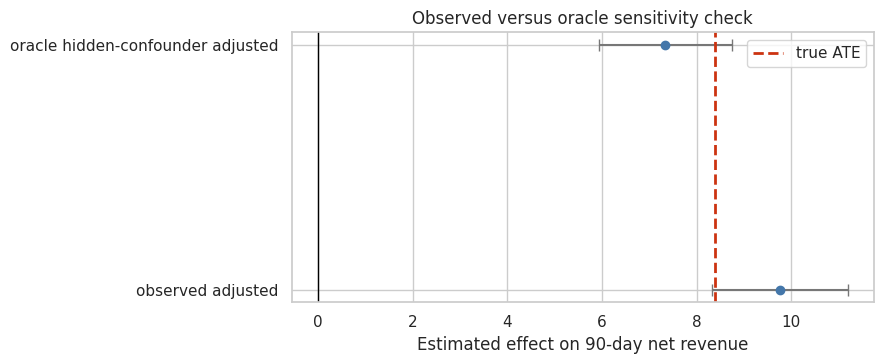
Discussion
The oracle comparison is a teaching device. In a real project we do not have executive_attention. If the oracle estimate differs from the observed-adjusted estimate, it shows why hidden confounding matters.
The placebo estimate is also important. Current treatment cannot cause prior-period revenue. A nonzero placebo effect is a warning about residual confounding or selection.
6. Benchmark Confounder Analysis
One practical sensitivity habit is to ask:
How much does the treatment coefficient move when important observed covariates are removed?
Those movements do not prove how large unobserved confounding is, but they give a scale for reasoning about omitted variables.
def refit_without_covariate(df, contract, omitted):
covariates = [var for var in contract['allowed_covariates'] if var != omitted]
categorical = [var for var in contract['categorical_covariates'] if var in covariates]
result = fit_effect_model(df, contract['outcome'], contract['treatment'], covariates, categorical)
return result['estimate']
benchmark_rows = []
for covariate in analysis_contract['allowed_covariates']:
reduced_estimate = refit_without_covariate(raw, analysis_contract, covariate)
shift = reduced_estimate - observed_result['estimate']
benchmark_rows.append(
{
'omitted_covariate': covariate,
'estimate_without_covariate': reduced_estimate,
'shift_from_full_model': shift,
'abs_shift': abs(shift),
'shift_as_share_of_estimate': abs(shift) / abs(observed_result['estimate']),
}
)
benchmark_table = pd.DataFrame(benchmark_rows).sort_values('abs_shift', ascending=False)
benchmark_table| omitted_covariate | estimate_without_covariate | shift_from_full_model | abs_shift | shift_as_share_of_estimate | |
|---|---|---|---|---|---|
| 7 | risk_score | 10.041754 | 0.283853 | 0.283853 | 0.029090 |
| 5 | contract_value | 10.013070 | 0.255169 | 0.255169 | 0.026150 |
| 6 | observed_health | 9.734774 | -0.023127 | 0.023127 | 0.002370 |
| 0 | segment | 9.776957 | 0.019056 | 0.019056 | 0.001953 |
| 4 | prior_support_tickets | 9.741525 | -0.016376 | 0.016376 | 0.001678 |
| 1 | region | 9.768545 | 0.010644 | 0.010644 | 0.001091 |
| 2 | tenure_months | 9.765030 | 0.007129 | 0.007129 | 0.000731 |
| 3 | baseline_usage | 9.752294 | -0.005607 | 0.005607 | 0.000575 |
fig, ax = plt.subplots(figsize=(8, 4.8))
sns.barplot(data=benchmark_table, y='omitted_covariate', x='shift_from_full_model', color='#AA3377', ax=ax)
ax.axvline(0, color='black', linewidth=1)
ax.set_title('Coefficient movement when observed covariates are omitted')
ax.set_xlabel('Shift in estimated treatment effect')
ax.set_ylabel('')
plt.tight_layout()
plt.show()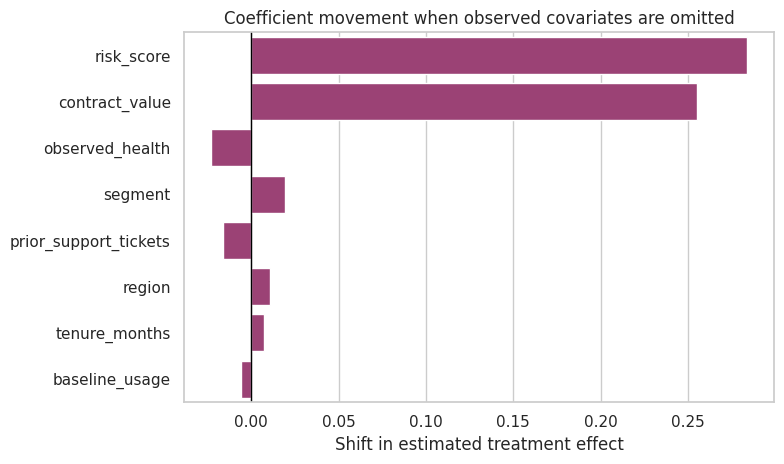
Discussion
If omitting one observed covariate moves the estimate substantially, then an unobserved covariate of comparable importance could also matter. Benchmarking makes that conversation concrete.
7. Residual-Confounding Sensitivity Grid
We now ask a hypothetical question:
Suppose there is one unobserved confounder left after adjusting for observed covariates. If that confounder has correlation
r_tuwith the residualized treatment andr_yuwith the residualized outcome, how would the treatment coefficient change?
This is a stylized sensitivity analysis. It is not a substitute for design knowledge, but it makes the tipping point visible.
def design_matrix_for_contract(df, covariates, categorical):
X = pd.get_dummies(df[covariates], columns=categorical, drop_first=True, dtype=float)
X = pd.concat([pd.Series(1.0, index=X.index, name='intercept'), X], axis=1)
return X
def residualize(series, X):
y = np.asarray(series, dtype=float)
beta = np.linalg.lstsq(np.asarray(X, dtype=float), y, rcond=None)[0]
return y - np.asarray(X, dtype=float) @ beta
def coefficient_with_unobserved_confounder(resid_t, resid_y, r_tu, r_yu):
resid_t = np.asarray(resid_t, dtype=float)
resid_y = np.asarray(resid_y, dtype=float)
var_t = np.var(resid_t, ddof=0)
var_y = np.var(resid_y, ddof=0)
cov_ty = np.mean((resid_t - resid_t.mean()) * (resid_y - resid_y.mean()))
sd_t = np.sqrt(var_t)
sd_y = np.sqrt(var_y)
cov_tu = r_tu * sd_t
cov_yu = r_yu * sd_y
denom = var_t - cov_tu ** 2
if denom <= 1e-12:
return np.nan
return (cov_ty - cov_tu * cov_yu) / denom
X_observed = design_matrix_for_contract(raw, analysis_contract['allowed_covariates'], analysis_contract['categorical_covariates'])
resid_t = residualize(raw[analysis_contract['treatment']], X_observed)
resid_y = residualize(raw[analysis_contract['outcome']], X_observed)
resid_placebo = residualize(raw[analysis_contract['placebo_outcome']], X_observed)
r_values = np.round(np.linspace(-0.60, 0.60, 49), 3)
grid_rows = []
for r_tu in r_values:
for r_yu in r_values:
adjusted_coef = coefficient_with_unobserved_confounder(resid_t, resid_y, r_tu, r_yu)
grid_rows.append({'r_tu': r_tu, 'r_yu': r_yu, 'sensitivity_adjusted_effect': adjusted_coef})
sensitivity_grid = pd.DataFrame(grid_rows)
sensitivity_grid.head()| r_tu | r_yu | sensitivity_adjusted_effect | |
|---|---|---|---|
| 0 | -0.6 | -0.600 | -14.525436 |
| 1 | -0.6 | -0.575 | -13.284929 |
| 2 | -0.6 | -0.550 | -12.044423 |
| 3 | -0.6 | -0.525 | -10.803916 |
| 4 | -0.6 | -0.500 | -9.563410 |
zero_crossing = sensitivity_grid.query('sensitivity_adjusted_effect <= 0').copy()
if zero_crossing.empty:
tipping_point = {'exists_in_grid': False, 'min_correlation_distance': np.nan, 'r_tu': np.nan, 'r_yu': np.nan, 'effect_at_tipping_point': np.nan}
else:
zero_crossing['correlation_distance'] = np.sqrt(zero_crossing['r_tu'] ** 2 + zero_crossing['r_yu'] ** 2)
row = zero_crossing.sort_values('correlation_distance').iloc[0]
tipping_point = {
'exists_in_grid': True,
'min_correlation_distance': float(row['correlation_distance']),
'r_tu': float(row['r_tu']),
'r_yu': float(row['r_yu']),
'effect_at_tipping_point': float(row['sensitivity_adjusted_effect']),
}
tipping_point{'exists_in_grid': True,
'min_correlation_distance': 0.6189709201569974,
'r_tu': 0.425,
'r_yu': 0.45,
'effect_at_tipping_point': -0.44501253597670876}heatmap_data = sensitivity_grid.pivot(index='r_yu', columns='r_tu', values='sensitivity_adjusted_effect')
fig, ax = plt.subplots(figsize=(9, 7))
sns.heatmap(heatmap_data, cmap='coolwarm', center=0, cbar_kws={'label': 'Adjusted effect'}, ax=ax)
ax.set_title('Residual-confounding sensitivity grid')
ax.set_xlabel('Correlation with residual treatment: r_tu')
ax.set_ylabel('Correlation with residual outcome: r_yu')
xticks = np.linspace(0, len(r_values) - 1, 7, dtype=int)
yticks = np.linspace(0, len(r_values) - 1, 7, dtype=int)
ax.set_xticks(xticks + 0.5)
ax.set_xticklabels([f'{r_values[i]:.2f}' for i in xticks], rotation=0)
ax.set_yticks(yticks + 0.5)
ax.set_yticklabels([f'{r_values[i]:.2f}' for i in yticks], rotation=0)
plt.tight_layout()
plt.show()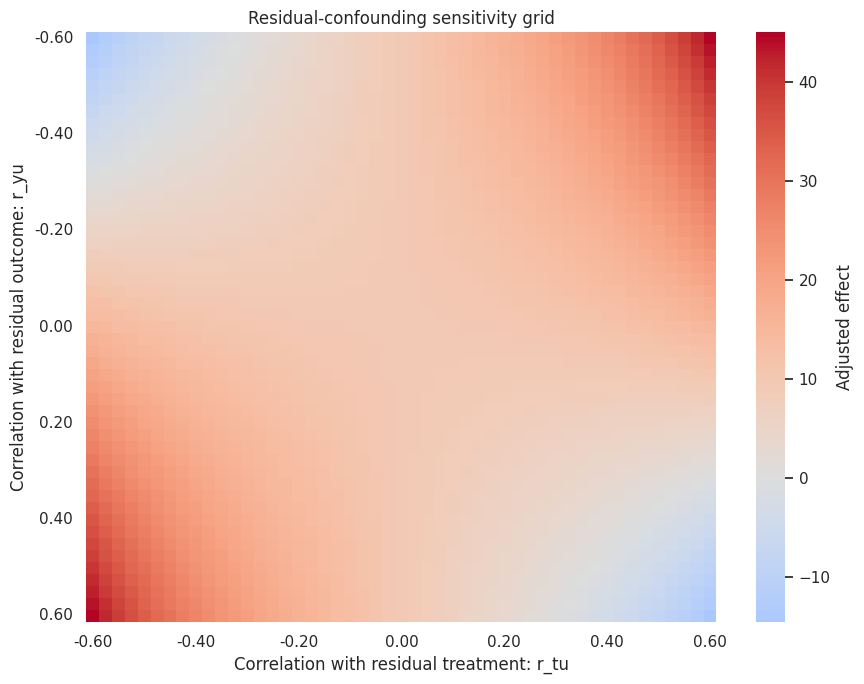
Discussion
The grid is not saying that a particular hidden confounder exists. It is saying how strong one would need to be, on the residual scale, to change the conclusion.
If the effect reverses under very small residual correlations, the design is fragile. If reversal requires implausibly strong residual correlations, the estimate is more robust to this specific form of hidden confounding. That still does not prove causality.
8. Placebo Outcome Sensitivity
A placebo outcome gives a different lens. Since treatment happens after prior-period revenue, any apparent effect on prior_period_revenue is evidence that treated and untreated accounts differ in ways the observed model has not fully removed.
placebo_summary = {
'placebo_outcome': analysis_contract['placebo_outcome'],
'placebo_estimate': placebo_result['estimate'],
'placebo_robust_se': placebo_result['robust_se'],
'placebo_ci_low': placebo_result['ci_low'],
'placebo_ci_high': placebo_result['ci_high'],
'absolute_placebo_estimate': abs(placebo_result['estimate']),
'placebo_as_share_of_observed_effect': abs(placebo_result['estimate']) / abs(observed_result['estimate']),
}
placebo_summary{'placebo_outcome': 'prior_period_revenue',
'placebo_estimate': 2.1943210894276772,
'placebo_robust_se': 0.6043799867184138,
'placebo_ci_low': 1.0097363154595862,
'placebo_ci_high': 3.3789058633957683,
'absolute_placebo_estimate': 2.1943210894276772,
'placebo_as_share_of_observed_effect': 0.2248763474696809}fig, ax = plt.subplots(figsize=(8, 3.6))
placebo_plot = pd.DataFrame(
[
{'estimand': 'main outcome estimate', 'estimate': observed_result['estimate'], 'ci_low': observed_result['ci_low'], 'ci_high': observed_result['ci_high']},
{'estimand': 'placebo prior outcome', 'estimate': placebo_result['estimate'], 'ci_low': placebo_result['ci_low'], 'ci_high': placebo_result['ci_high']},
]
)
ax.errorbar(
placebo_plot['estimate'],
placebo_plot['estimand'],
xerr=[placebo_plot['estimate'] - placebo_plot['ci_low'], placebo_plot['ci_high'] - placebo_plot['estimate']],
fmt='o',
color='#228833',
ecolor='#777777',
capsize=4,
)
ax.axvline(0, color='black', linewidth=1)
ax.set_title('Main estimate versus placebo outcome estimate')
ax.set_xlabel('Estimated coefficient on onboarding_specialist')
ax.set_ylabel('')
plt.tight_layout()
plt.show()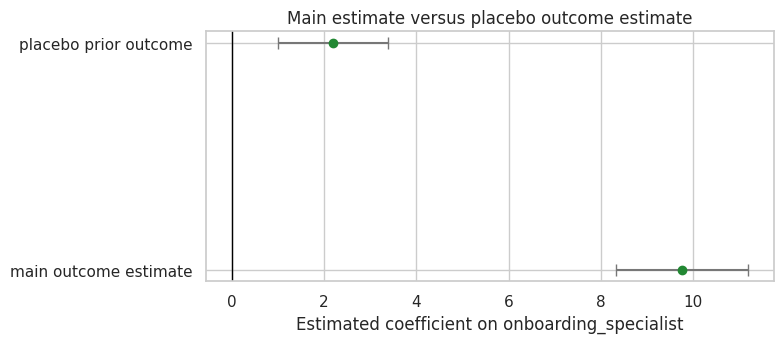
Discussion
A placebo failure does not tell us the exact bias in the main outcome. It tells us the design can produce an apparent treatment effect where no treatment effect should exist. That should change how cautiously we communicate the main estimate.
9. Sensitivity Gates
We now turn the sensitivity evidence into a decision recommendation. The gates are deliberately conservative.
def classify_sensitivity_gates(observed_result, oracle_result, benchmark_table, placebo_summary, tipping_point, contract):
thresholds = contract['diagnostic_thresholds']
max_benchmark_shift = benchmark_table['abs_shift'].max()
oracle_shift = abs(observed_result['estimate'] - oracle_result['estimate'])
gates = [
{
'gate': 'oracle shift in synthetic lab',
'metric': 'abs observed-minus-oracle estimate',
'value': oracle_shift,
'status': 'caution' if oracle_shift > max_benchmark_shift else 'pass',
'interpretation': 'How much the estimate changes when hidden confounder is included. Teaching-only diagnostic.',
},
{
'gate': 'largest observed-covariate benchmark',
'metric': 'max single-covariate shift',
'value': max_benchmark_shift,
'status': 'caution' if max_benchmark_shift / abs(observed_result['estimate']) > thresholds['benchmark_shift_caution_ratio'] else 'pass',
'interpretation': 'Scale of coefficient movement from omitting observed covariates.',
},
{
'gate': 'placebo outcome',
'metric': 'absolute placebo estimate',
'value': placebo_summary['absolute_placebo_estimate'],
'status': 'caution' if placebo_summary['absolute_placebo_estimate'] > thresholds['placebo_caution_revenue_units'] else 'pass',
'interpretation': 'Apparent effect on pre-treatment outcome.',
},
{
'gate': 'tipping point proximity',
'metric': 'minimum residual-correlation distance to sign reversal',
'value': tipping_point['min_correlation_distance'],
'status': 'caution' if tipping_point['exists_in_grid'] and tipping_point['min_correlation_distance'] < thresholds['tipping_point_distance_caution'] else 'pass',
'interpretation': 'How close the estimate is to sign reversal under residual confounding grid.',
},
]
gate_table = pd.DataFrame(gates)
if (gate_table['status'] == 'caution').sum() >= 2:
recommendation = 'collect_more_evidence_before_strong_causal_claim'
elif (gate_table['status'] == 'caution').sum() == 1:
recommendation = 'report_with_caution_and_sensitivity_limitations'
else:
recommendation = 'sensitivity_checks_support_cautious_reporting'
return gate_table, recommendation
sensitivity_gate_table, sensitivity_recommendation = classify_sensitivity_gates(
observed_result,
oracle_result,
benchmark_table,
placebo_summary,
tipping_point,
analysis_contract,
)
sensitivity_gate_table| gate | metric | value | status | interpretation | |
|---|---|---|---|---|---|
| 0 | oracle shift in synthetic lab | abs observed-minus-oracle estimate | 2.421653 | caution | How much the estimate changes when hidden confounder is included. Teaching-only diagnostic. |
| 1 | largest observed-covariate benchmark | max single-covariate shift | 0.283853 | pass | Scale of coefficient movement from omitting observed covariates. |
| 2 | placebo outcome | absolute placebo estimate | 2.194321 | pass | Apparent effect on pre-treatment outcome. |
| 3 | tipping point proximity | minimum residual-correlation distance to sign reversal | 0.618971 | pass | How close the estimate is to sign reversal under residual confounding grid. |
sensitivity_recommendation'report_with_caution_and_sensitivity_limitations'Discussion
The gate recommendation is not a universal rule. It is a reporting discipline. It forces the analyst to say when sensitivity evidence weakens the causal story.
10. Build a Sensitivity Report Bundle
We now assemble the sensitivity evidence into a machine-readable report for AI review.
def dataframe_records(table, max_rows=30):
return json.loads(table.head(max_rows).to_json(orient='records', date_format='iso'))
sensitivity_report = {
'analysis_contract': analysis_contract,
'truth_available_only_because_synthetic': truth,
'observed_adjusted_estimate': {k: v for k, v in observed_result.items() if k != 'model'},
'oracle_hidden_confounder_estimate': {k: v for k, v in oracle_result.items() if k != 'model'},
'placebo_estimate': {k: v for k, v in placebo_result.items() if k != 'model'},
'benchmark_covariate_table': dataframe_records(benchmark_table, max_rows=12),
'tipping_point': tipping_point,
'sensitivity_grid_summary': {
'grid_min_effect': float(sensitivity_grid['sensitivity_adjusted_effect'].min()),
'grid_max_effect': float(sensitivity_grid['sensitivity_adjusted_effect'].max()),
'share_grid_effect_below_zero': float((sensitivity_grid['sensitivity_adjusted_effect'] <= 0).mean()),
},
'placebo_summary': placebo_summary,
'sensitivity_gates': dataframe_records(sensitivity_gate_table, max_rows=10),
'sensitivity_recommendation': sensitivity_recommendation,
'explicit_warnings': [
'Sensitivity analysis does not prove the effect is causal.',
'The oracle estimate uses a hidden confounder that exists only in this synthetic teaching example.',
'A placebo outcome warning should reduce confidence even if the main estimate is statistically significant.',
'LLM summaries should not convert sensitivity checks into a robustness certificate.',
],
}
print(json.dumps(sensitivity_report, indent=2)[:5200]){
"analysis_contract": {
"analysis_id": "onboarding_specialist_sensitivity_v1",
"estimand": "ATE of onboarding_specialist on net_revenue_90d among eligible accounts",
"treatment": "onboarding_specialist",
"outcome": "net_revenue_90d",
"placebo_outcome": "prior_period_revenue",
"allowed_covariates": [
"segment",
"region",
"tenure_months",
"baseline_usage",
"prior_support_tickets",
"contract_value",
"observed_health",
"risk_score"
],
"categorical_covariates": [
"segment",
"region"
],
"oracle_hidden_confounders": [
"executive_attention"
],
"forbidden_variables": [
"product_adoption_30d",
"expansion_pipeline_120d",
"true_effect"
],
"diagnostic_thresholds": {
"placebo_caution_revenue_units": 4.0,
"tipping_point_distance_caution": 0.35,
"benchmark_shift_caution_ratio": 1.0
}
},
"truth_available_only_because_synthetic": {
"true_ate": 8.388275446972925,
"true_att": 8.50067894836023,
"treatment_rate": 0.685,
"hidden_strength": 0.65
},
"observed_adjusted_estimate": {
"formula": "net_revenue_90d ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_value + observed_health + risk_score",
"estimate": 9.757900793561795,
"robust_se": 0.729625671578333,
"ci_low": 8.327834477268262,
"ci_high": 11.187967109855329,
"p_value": 8.597863559654592e-41,
"rows": 5200
},
"oracle_hidden_confounder_estimate": {
"formula": "net_revenue_90d ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_value + observed_health + risk_score + executive_attention",
"estimate": 7.336248179580361,
"robust_se": 0.7170837832752662,
"ci_low": 5.930763964360839,
"ci_high": 8.741732394799882,
"p_value": 1.4451474851989958e-24,
"rows": 5200
},
"placebo_estimate": {
"formula": "prior_period_revenue ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_value + observed_health + risk_score",
"estimate": 2.1943210894276772,
"robust_se": 0.6043799867184138,
"ci_low": 1.0097363154595862,
"ci_high": 3.3789058633957683,
"p_value": 0.0002826559845974888,
"rows": 5200
},
"benchmark_covariate_table": [
{
"omitted_covariate": "risk_score",
"estimate_without_covariate": 10.041754266,
"shift_from_full_model": 0.2838534725,
"abs_shift": 0.2838534725,
"shift_as_share_of_estimate": 0.0290896043
},
{
"omitted_covariate": "contract_value",
"estimate_without_covariate": 10.013070119,
"shift_from_full_model": 0.2551693254,
"abs_shift": 0.2551693254,
"shift_as_share_of_estimate": 0.0261500225
},
{
"omitted_covariate": "observed_health",
"estimate_without_covariate": 9.734774011,
"shift_from_full_model": -0.0231267825,
"abs_shift": 0.0231267825,
"shift_as_share_of_estimate": 0.0023700571
},
{
"omitted_covariate": "segment",
"estimate_without_covariate": 9.7769572867,
"shift_from_full_model": 0.0190564932,
"abs_shift": 0.0190564932,
"shift_as_share_of_estimate": 0.0019529296
},
{
"omitted_covariate": "prior_support_tickets",
"estimate_without_covariate": 9.7415246054,
"shift_from_full_model": -0.0163761881,
"abs_shift": 0.0163761881,
"shift_as_share_of_estimate": 0.0016782491
},
{
"omitted_covariate": "region",
"estimate_without_covariate": 9.7685447453,
"shift_from_full_model": 0.0106439518,
"abs_shift": 0.0106439518,
"shift_as_share_of_estimate": 0.0010908034
},
{
"omitted_covariate": "tenure_months",
"estimate_without_covariate": 9.765030247,
"shift_from_full_model": 0.0071294534,
"abs_shift": 0.0071294534,
"shift_as_share_of_estimate": 0.0007306339
},
{
"omitted_covariate": "baseline_usage",
"estimate_without_covariate": 9.7522938182,
"shift_from_full_model": -0.0056069753,
"abs_shift": 0.0056069753,
"shift_as_share_of_estimate": 0.0005746088
}
],
"tipping_point": {
"exists_in_grid": true,
"min_correlation_distance": 0.6189709201569974,
"r_tu": 0.425,
"r_yu": 0.45,
"effect_at_tipping_point": -0.44501253597670876
},
"sensitivity_grid_summary": {
"grid_min_effect": -14.525435716611746,
"grid_max_effect": 45.01887569649227,
"share_grid_effect_below_zero": 0.07913369429404415
},
"placebo_summary": {
"placebo_outcome": "prior_period_revenue",
"placebo_estimate": 2.1943210894276772,
"placebo_robust_se": 0.6043799867184138,
"placebo_ci_low": 1.0097363154595862,
"placebo_ci_high": 3.3789058633957683,
"absolute_placebo_estimate": 2.1943210894276772,
"placebo_as_share_of_observed_effect": 0.2248763474696809
},
"sensitivity_gates": [
{
"gate": "oracle shift in synthetic lab",
"metric": "abs observed-minus-oracle estimate",
"Discussion
The report bundle contains numbers, gates, and warnings. This is the right object for an AI assistant to summarize. It is far safer than asking, “Is my result robust?”
12. Structured AI Sensitivity Review
The model will summarize the sensitivity report. It must not declare the result “robust” unless the gates support that language, and even then it must preserve limitations.
class SensitivityReview(BaseModel):
analysis_summary: str = Field(description='Brief summary of estimand and design.')
main_estimate_interpretation: list[str] = Field(description='How to interpret the main adjusted estimate.')
benchmark_interpretation: list[str] = Field(description='Interpretation of observed-covariate benchmark shifts.')
placebo_interpretation: list[str] = Field(description='Interpretation of placebo outcome evidence.')
tipping_point_interpretation: list[str] = Field(description='Interpretation of residual-confounding tipping point.')
recommendation: Literal['cautious_reporting', 'collect_more_evidence', 'do_not_make_causal_claim'] = Field(description='Recommended communication posture.')
recommendation_rationale: list[str] = Field(description='Reasons for the recommendation.')
additional_sensitivity_checks: list[str] = Field(description='Additional checks to run next.')
communication_warnings: list[str] = Field(description='Warnings for stakeholder-facing language.')
stakeholder_summary: str = Field(description='Plain-language summary for decision makers.')
confidence: Literal['low', 'medium', 'high'] = Field(description='Confidence in the review given the report.')
REVIEW_SCALAR_FIELDS = ['analysis_summary', 'recommendation', 'stakeholder_summary', 'confidence']
REVIEW_LIST_FIELDS = [
'main_estimate_interpretation',
'benchmark_interpretation',
'placebo_interpretation',
'tipping_point_interpretation',
'recommendation_rationale',
'additional_sensitivity_checks',
'communication_warnings',
]
REVIEW_VALUE_ALIASES = {
'recommendation': {
'cautious_reporting': 'cautious_reporting',
'cautiousreporting': 'cautious_reporting',
'cautious': 'cautious_reporting',
'caution': 'cautious_reporting',
'report_cautiously': 'cautious_reporting',
'proceed_with_caution': 'cautious_reporting',
'collect_more_evidence': 'collect_more_evidence',
'collectmoreevidence': 'collect_more_evidence',
'more_evidence': 'collect_more_evidence',
'gather_more_evidence': 'collect_more_evidence',
'additional_evidence': 'collect_more_evidence',
'do_not_make_causal_claim': 'do_not_make_causal_claim',
'donotmakecausalclaim': 'do_not_make_causal_claim',
'do_not_claim_causality': 'do_not_make_causal_claim',
'no_causal_claim': 'do_not_make_causal_claim',
'avoid_causal_claim': 'do_not_make_causal_claim',
},
'confidence': {
'low': 'low',
'medium': 'medium',
'moderate': 'medium',
'high': 'high',
},
}
REVIEW_DEFAULTS = {
'analysis_summary': '',
'main_estimate_interpretation': [],
'benchmark_interpretation': [],
'placebo_interpretation': [],
'tipping_point_interpretation': [],
'recommendation_rationale': [],
'additional_sensitivity_checks': [],
'communication_warnings': [],
'stakeholder_summary': '',
'confidence': 'medium',
}
REVIEW_ALIASES = {
'summary': 'analysis_summary',
'main_estimate': 'main_estimate_interpretation',
'benchmark': 'benchmark_interpretation',
'placebo': 'placebo_interpretation',
'tipping_point': 'tipping_point_interpretation',
'rationale': 'recommendation_rationale',
'next_checks': 'additional_sensitivity_checks',
'warnings': 'communication_warnings',
}
def parse_sensitivity_review(raw_output):
result = parse_pydantic_output(
raw_output,
SensitivityReview,
scalar_fields=REVIEW_SCALAR_FIELDS,
list_fields=REVIEW_LIST_FIELDS,
field_aliases=REVIEW_ALIASES,
value_aliases=REVIEW_VALUE_ALIASES,
defaults=REVIEW_DEFAULTS,
)
return result.parsed, result.json_text, result.notesSYSTEM_REVIEW_MESSAGE = (
'You are a careful causal inference sensitivity-analysis reviewer.\n'
'Rules:\n'
'- Use only the provided sensitivity report.\n'
'- Return valid JSON only. No markdown. No preamble.\n'
'- Do not claim sensitivity analysis proves causality.\n'
'- Mention benchmark confounders, placebo outcome evidence, tipping point, and oracle limitations when available.\n'
'- Do not treat synthetic oracle columns as available in real data.\n'
'- If placebo or tipping-point gates are cautionary, do not recommend strong causal claims.\n'
'- Mention brittleness and rerun/model-output instability where appropriate.'
)
def build_review_prompt(report):
schema_hint = {
'analysis_summary': 'string',
'main_estimate_interpretation': ['string'],
'benchmark_interpretation': ['string'],
'placebo_interpretation': ['string'],
'tipping_point_interpretation': ['string'],
'recommendation': 'cautious_reporting | collect_more_evidence | do_not_make_causal_claim',
'recommendation_rationale': ['string'],
'additional_sensitivity_checks': ['string'],
'communication_warnings': ['string'],
'stakeholder_summary': 'string',
'confidence': 'low | medium | high',
}
return textwrap.dedent(
f'''
Produce a SensitivityReview JSON object using this schema.
Use the exact enum strings shown in the schema. Every field is required; use [] when no useful item is available.
Return one JSON object only.
{json.dumps(schema_hint, indent=2)}
Sensitivity report:
{json.dumps(report, indent=2)}
'''
).strip()
review_prompt = build_review_prompt(sensitivity_report)
print(review_prompt[:3000])Produce a SensitivityReview JSON object using this schema.
Use the exact enum strings shown in the schema. Every field is required; use [] when no useful item is available.
Return one JSON object only.
{
"analysis_summary": "string",
"main_estimate_interpretation": [
"string"
],
"benchmark_interpretation": [
"string"
],
"placebo_interpretation": [
"string"
],
"tipping_point_interpretation": [
"string"
],
"recommendation": "cautious_reporting | collect_more_evidence | do_not_make_causal_claim",
"recommendation_rationale": [
"string"
],
"additional_sensitivity_checks": [
"string"
],
"communication_warnings": [
"string"
],
"stakeholder_summary": "string",
"confidence": "low | medium | high"
}
Sensitivity report:
{
"analysis_contract": {
"analysis_id": "onboarding_specialist_sensitivity_v1",
"estimand": "ATE of onboarding_specialist on net_revenue_90d among eligible accounts",
"treatment": "onboarding_specialist",
"outcome": "net_revenue_90d",
"placebo_outcome": "prior_period_revenue",
"allowed_covariates": [
"segment",
"region",
"tenure_months",
"baseline_usage",
"prior_support_tickets",
"contract_value",
"observed_health",
"risk_score"
],
"categorical_covariates": [
"segment",
"region"
],
"oracle_hidden_confounders": [
"executive_attention"
],
"forbidden_variables": [
"product_adoption_30d",
"expansion_pipeline_120d",
"true_effect"
],
"diagnostic_thresholds": {
"placebo_caution_revenue_units": 4.0,
"tipping_point_distance_caution": 0.35,
"benchmark_shift_caution_ratio": 1.0
}
},
"truth_available_only_because_synthetic": {
"true_ate": 8.388275446972925,
"true_att": 8.50067894836023,
"treatment_rate": 0.685,
"hidden_strength": 0.65
},
"observed_adjusted_estimate": {
"formula": "net_revenue_90d ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_value + observed_health + risk_score",
"estimate": 9.757900793561795,
"robust_se": 0.729625671578333,
"ci_low": 8.327834477268262,
"ci_high": 11.187967109855329,
"p_value": 8.597863559654592e-41,
"rows": 5200
},
"oracle_hidden_confounder_estimate": {
"formula": "net_revenue_90d ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_value + observed_health + risk_score + executive_attention",
"estimate": 7.336248179580361,
"robust_se": 0.7170837832752662,
"ci_low": 5.930763964360839,
"ci_high": 8.741732394799882,
"p_value": 1.4451474851989958e-24,
"rows": 5200
},
"placebo_estimate": {
"formula": "prior_period_revenue ~ onboarding_specialist + C(segment) + C(region) + tenure_months + baseline_usage + prior_support_tickets + contract_13. Run One Local Model
We start with the fast local model. A good answer should summarize the sensitivity evidence without turning it into a robustness certificate.
if RUN_LIVE_LOCAL_LLM:
raw_review_output = local_chat(
review_prompt,
system_message=SYSTEM_REVIEW_MESSAGE,
model_id=MODEL_ID,
max_new_tokens=MAX_NEW_TOKENS,
temperature=TEMPERATURE,
)
display(Markdown('### Raw model output'))
display(Markdown(f'```json\n{clean_generated_text(raw_review_output)}\n```'))
else:
raw_review_output = ''
print('RUN_LIVE_LOCAL_LLM is False. Skipping live local LLM call in the rendered notebook.')Raw model output
{
"analysis_summary": "The analysis evaluates the ATE of onboarding specialists on net revenue 90 days post-treatment, adjusting for several covariates. The observed adjusted estimate suggests a positive effect, but sensitivity analyses indicate potential confounding.",
"main_estimate_interpretation": [
"The observed adjusted estimate indicates a positive effect of onboarding specialists on net revenue 90 days post-treatment."
],
"benchmark_interpretation": [
"Omitting 'risk_score' and 'contract_value' results in the largest shifts in the estimate, but these shifts are relatively small compared to the overall estimate."
],
"placebo_interpretation": [
"The placebo outcome shows a non-zero effect, which is a cautionary signal as it should be close to zero if there were no confounding."
],
"tipping_point_interpretation": [
"The tipping point is relatively far from the current estimate, suggesting some robustness to unmeasured confounding."
],
"recommendation": "cautious_reporting",
"recommendation_rationale": [
"The placebo outcome shows a non-zero effect, indicating potential confounding issues.",
"The oracle shift in synthetic lab is at the caution threshold, suggesting the estimate could change significantly with additional confounding."
],
"additional_sensitivity_checks": [
"Conduct further sensitivity analyses with different sets of covariates.",
"Explore the impact of interaction terms between covariates."
],
"communication_warnings": [
"Avoid making strong causal claims based solely on this analysis.",
"Emphasize the limitations of the sensitivity analysis and the need for further validation."
],
"stakeholder_summary": "The analysis suggests a positive effect of onboarding specialists on net revenue, but sensitivity checks indicate potential confounding. Further investigation is recommended before making definitive causal claims.",
"confidence": "medium"
}if raw_review_output:
try:
sensitivity_review, sensitivity_review_json, review_parser_notes = parse_sensitivity_review(raw_review_output)
single_model_parse_error = ''
display(Markdown('### Parsed sensitivity review'))
display(Markdown(f'```json\n{sensitivity_review.model_dump_json(indent=2)}\n```'))
print('Parser notes:', review_parser_notes)
except Exception as error:
sensitivity_review = None
sensitivity_review_json = ''
review_parser_notes = []
single_model_parse_error = clean_generated_text(repr(error))
print('The single-model output could not be parsed. This is a useful brittleness signal, not a reason to trust the output silently.')
print(single_model_parse_error[:1200])
else:
sensitivity_review = None
sensitivity_review_json = ''
review_parser_notes = []
single_model_parse_error = ''Parsed sensitivity review
{
"analysis_summary": "The analysis evaluates the ATE of onboarding specialists on net revenue 90 days post-treatment, adjusting for several covariates. The observed adjusted estimate suggests a positive effect, but sensitivity analyses indicate potential confounding.",
"main_estimate_interpretation": [
"The observed adjusted estimate indicates a positive effect of onboarding specialists on net revenue 90 days post-treatment."
],
"benchmark_interpretation": [
"Omitting 'risk_score' and 'contract_value' results in the largest shifts in the estimate, but these shifts are relatively small compared to the overall estimate."
],
"placebo_interpretation": [
"The placebo outcome shows a non-zero effect, which is a cautionary signal as it should be close to zero if there were no confounding."
],
"tipping_point_interpretation": [
"The tipping point is relatively far from the current estimate, suggesting some robustness to unmeasured confounding."
],
"recommendation": "cautious_reporting",
"recommendation_rationale": [
"The placebo outcome shows a non-zero effect, indicating potential confounding issues.",
"The oracle shift in synthetic lab is at the caution threshold, suggesting the estimate could change significantly with additional confounding."
],
"additional_sensitivity_checks": [
"Conduct further sensitivity analyses with different sets of covariates.",
"Explore the impact of interaction terms between covariates."
],
"communication_warnings": [
"Avoid making strong causal claims based solely on this analysis.",
"Emphasize the limitations of the sensitivity analysis and the need for further validation."
],
"stakeholder_summary": "The analysis suggests a positive effect of onboarding specialists on net revenue, but sensitivity checks indicate potential confounding. Further investigation is recommended before making definitive causal claims.",
"confidence": "medium"
}Parser notes: []14. Audit the AI Sensitivity Review
A good sensitivity review must mention the actual sensitivity evidence and avoid causal overclaiming.
def review_text(review):
if review is None:
return ''
return clean_generated_text(json.dumps(review.model_dump(), sort_keys=True)).lower()
def contains_any(text, terms):
return any(term.lower() in text for term in terms)
def expected_recommendation_from_sensitivity(gates):
cautions = (gates['status'] == 'caution').sum()
if cautions >= 2:
return 'collect_more_evidence'
if cautions == 1:
return 'cautious_reporting'
return 'cautious_reporting'
expected_recommendation = expected_recommendation_from_sensitivity(sensitivity_gate_table)
def recommendation_not_less_cautious(model_rec, expected_rec):
order = {'cautious_reporting': 0, 'collect_more_evidence': 1, 'do_not_make_causal_claim': 2}
return order.get(model_rec, -1) >= order.get(expected_rec, 0)
def score_sensitivity_review(review, expected_recommendation):
text = review_text(review)
recommendation = review.recommendation if review is not None else ''
checks = {
'states estimand': analysis_contract['treatment'] in text and analysis_contract['outcome'] in text,
'mentions benchmark confounders': contains_any(text, ['benchmark', 'observed covariate', 'omitted covariate']),
'mentions placebo outcome': contains_any(text, ['placebo', analysis_contract['placebo_outcome']]),
'mentions tipping point': contains_any(text, ['tipping', 'residual confounding', 'sign reversal']),
'mentions oracle limitation': contains_any(text, ['oracle', 'synthetic', 'hidden confounder', 'not available in real data']),
'recommendation is not less cautious than gates': recommendation_not_less_cautious(recommendation, expected_recommendation),
'does not claim sensitivity proves causality': not contains_any(text, ['proves causality', 'proves the effect', 'certifies robustness', 'eliminates unmeasured confounding']),
'includes communication warnings': len(review.communication_warnings) >= 2 if review is not None else False,
'includes additional checks': len(review.additional_sensitivity_checks) >= 2 if review is not None else False,
'mentions brittleness or caution': contains_any(text, ['caution', 'brittle', 'sensitivity', 'fragile', 'uncertain']),
}
audit = pd.DataFrame([{'check': key, 'passed': bool(value)} for key, value in checks.items()])
audit['credit'] = audit['passed'].astype(int)
return audit, int(audit['credit'].sum()), len(audit)
if sensitivity_review is not None:
single_audit, single_score, single_max_score = score_sensitivity_review(sensitivity_review, expected_recommendation)
print(f'Expected recommendation from gates: {expected_recommendation}')
print(f'Score: {single_score}/{single_max_score}')
display(single_audit)
else:
print('No sensitivity review to audit because live model execution was skipped or parsing failed.')Expected recommendation from gates: cautious_reporting
Score: 9/10| check | passed | credit | |
|---|---|---|---|
| 0 | states estimand | False | 0 |
| 1 | mentions benchmark confounders | True | 1 |
| 2 | mentions placebo outcome | True | 1 |
| 3 | mentions tipping point | True | 1 |
| 4 | mentions oracle limitation | True | 1 |
| 5 | recommendation is not less cautious than gates | True | 1 |
| 6 | does not claim sensitivity proves causality | True | 1 |
| 7 | includes communication warnings | True | 1 |
| 8 | includes additional checks | True | 1 |
| 9 | mentions brittleness or caution | True | 1 |
Discussion
The model may be more conservative than the deterministic gates. That is acceptable. It should not be less conservative, and it should never say the sensitivity analysis proves the causal claim.
15. Optional All-Model Comparison
We now compare local models on two sensitivity reports: a moderate-hidden-confounding case and a strong-hidden-confounding case.
The full single-model review above uses a richer narrative schema. For the all-model comparison, we intentionally use a compact decision schema: key risk signals, recommendation, rationale, next checks, warnings, and confidence. This makes the comparison about causal judgment and schema reliability, not about whether every model can produce a long report with many fields.
The exact ranking can change across reruns and environments. That is expected. The durable lesson is that sensitivity-review assistants need to be evaluated against the numeric gates, not their confidence of tone. Remaining schema failures are useful evidence about which models are not reliable enough for automated causal-review workflows without additional guardrails.
def build_sensitivity_report_for_scenario(case_id, hidden_strength, overlap_noise, seed):
scenario_df, scenario_truth = simulate_onboarding_data(seed=seed, hidden_strength=hidden_strength, overlap_noise=overlap_noise)
obs = fit_effect_model(
scenario_df,
analysis_contract['outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'],
analysis_contract['categorical_covariates'],
)
oracle = fit_effect_model(
scenario_df,
analysis_contract['outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'] + analysis_contract['oracle_hidden_confounders'],
analysis_contract['categorical_covariates'],
)
placebo = fit_effect_model(
scenario_df,
analysis_contract['placebo_outcome'],
analysis_contract['treatment'],
analysis_contract['allowed_covariates'],
analysis_contract['categorical_covariates'],
)
bench_rows = []
for covariate in analysis_contract['allowed_covariates']:
covariates = [var for var in analysis_contract['allowed_covariates'] if var != covariate]
categorical = [var for var in analysis_contract['categorical_covariates'] if var in covariates]
reduced = fit_effect_model(scenario_df, analysis_contract['outcome'], analysis_contract['treatment'], covariates, categorical)
shift = reduced['estimate'] - obs['estimate']
bench_rows.append({'omitted_covariate': covariate, 'estimate_without_covariate': reduced['estimate'], 'shift_from_full_model': shift, 'abs_shift': abs(shift), 'shift_as_share_of_estimate': abs(shift) / abs(obs['estimate'])})
bench = pd.DataFrame(bench_rows).sort_values('abs_shift', ascending=False)
X = design_matrix_for_contract(scenario_df, analysis_contract['allowed_covariates'], analysis_contract['categorical_covariates'])
rt = residualize(scenario_df[analysis_contract['treatment']], X)
ry = residualize(scenario_df[analysis_contract['outcome']], X)
rows = []
for r_tu in r_values:
for r_yu in r_values:
rows.append({'r_tu': r_tu, 'r_yu': r_yu, 'sensitivity_adjusted_effect': coefficient_with_unobserved_confounder(rt, ry, r_tu, r_yu)})
grid = pd.DataFrame(rows)
zero = grid.query('sensitivity_adjusted_effect <= 0').copy()
if zero.empty:
tip = {'exists_in_grid': False, 'min_correlation_distance': np.nan, 'r_tu': np.nan, 'r_yu': np.nan, 'effect_at_tipping_point': np.nan}
else:
zero['correlation_distance'] = np.sqrt(zero['r_tu'] ** 2 + zero['r_yu'] ** 2)
row = zero.sort_values('correlation_distance').iloc[0]
tip = {'exists_in_grid': True, 'min_correlation_distance': float(row['correlation_distance']), 'r_tu': float(row['r_tu']), 'r_yu': float(row['r_yu']), 'effect_at_tipping_point': float(row['sensitivity_adjusted_effect'])}
placebo_sum = {
'placebo_outcome': analysis_contract['placebo_outcome'],
'placebo_estimate': placebo['estimate'],
'placebo_robust_se': placebo['robust_se'],
'placebo_ci_low': placebo['ci_low'],
'placebo_ci_high': placebo['ci_high'],
'absolute_placebo_estimate': abs(placebo['estimate']),
'placebo_as_share_of_observed_effect': abs(placebo['estimate']) / abs(obs['estimate']),
}
gates, rec = classify_sensitivity_gates(obs, oracle, bench, placebo_sum, tip, analysis_contract)
report = {
'case_id': case_id,
'analysis_contract': analysis_contract,
'truth_available_only_because_synthetic': scenario_truth,
'observed_adjusted_estimate': {k: v for k, v in obs.items() if k != 'model'},
'oracle_hidden_confounder_estimate': {k: v for k, v in oracle.items() if k != 'model'},
'placebo_estimate': {k: v for k, v in placebo.items() if k != 'model'},
'benchmark_covariate_table': dataframe_records(bench, max_rows=12),
'tipping_point': tip,
'sensitivity_grid_summary': {
'grid_min_effect': float(grid['sensitivity_adjusted_effect'].min()),
'grid_max_effect': float(grid['sensitivity_adjusted_effect'].max()),
'share_grid_effect_below_zero': float((grid['sensitivity_adjusted_effect'] <= 0).mean()),
},
'placebo_summary': placebo_sum,
'sensitivity_gates': dataframe_records(gates, max_rows=10),
'sensitivity_recommendation': rec,
'explicit_warnings': sensitivity_report['explicit_warnings'],
}
return report, expected_recommendation_from_sensitivity(gates)
moderate_report, moderate_expected = build_sensitivity_report_for_scenario('moderate_hidden_confounding', 0.35, 0.90, SEED + 1)
strong_report, strong_expected = build_sensitivity_report_for_scenario('strong_hidden_confounding', 1.05, 0.55, SEED + 2)
SENSITIVITY_EVAL_CASES = [
{'case_id': 'moderate_hidden_confounding', 'report': moderate_report, 'expected_recommendation': moderate_expected},
{'case_id': 'strong_hidden_confounding', 'report': strong_report, 'expected_recommendation': strong_expected},
]
[(case['case_id'], case['expected_recommendation']) for case in SENSITIVITY_EVAL_CASES][('moderate_hidden_confounding', 'cautious_reporting'),
('strong_hidden_confounding', 'collect_more_evidence')]SUMMARY_COLUMNS = [
'label', 'model_id', 'role', 'cases', 'schema_valid_cases', 'schema_repaired_cases',
'schema_reliability', 'mean_review_score', 'failure_types'
]
CASE_RESULT_COLUMNS = [
'label', 'model_id', 'role', 'case_id', 'expected_recommendation', 'model_recommendation', 'status',
'schema_valid', 'repair_used', 'repair_stage', 'error_type', 'review_score', 'max_review_score',
'review_score_share', 'error', 'raw_output_preview'
]
class SensitivityDecision(BaseModel):
analysis_summary: str = Field(description='One-sentence summary of the sensitivity-analysis situation.')
key_risk_signals: list[str] = Field(description='Most important sensitivity, placebo, benchmark, or tipping-point signals.')
recommendation: Literal['cautious_reporting', 'collect_more_evidence', 'do_not_make_causal_claim'] = Field(description='Recommended communication posture.')
recommendation_rationale: list[str] = Field(description='Reasons for the recommendation.')
additional_sensitivity_checks: list[str] = Field(description='Additional checks to run next.')
communication_warnings: list[str] = Field(description='Warnings for stakeholder-facing language.')
confidence: Literal['low', 'medium', 'high'] = Field(description='Confidence in the decision given the report.')
DECISION_SCALAR_FIELDS = ['analysis_summary', 'recommendation', 'confidence']
DECISION_LIST_FIELDS = [
'key_risk_signals',
'recommendation_rationale',
'additional_sensitivity_checks',
'communication_warnings',
]
DECISION_ALIASES = {
'summary': 'analysis_summary',
'risk_signals': 'key_risk_signals',
'risks': 'key_risk_signals',
'signals': 'key_risk_signals',
'rationale': 'recommendation_rationale',
'reasons': 'recommendation_rationale',
'next_checks': 'additional_sensitivity_checks',
'checks': 'additional_sensitivity_checks',
'warnings': 'communication_warnings',
}
DECISION_DEFAULTS = {
'analysis_summary': '',
'key_risk_signals': [],
'recommendation_rationale': [],
'additional_sensitivity_checks': [],
'communication_warnings': [],
'confidence': 'medium',
}
DECISION_REPAIR_PROMPT_TEMPLATE = textwrap.dedent(
'''
Your previous answer could not be parsed as the required SensitivityDecision JSON schema.
Convert the previous answer into valid JSON only. Do not add new causal claims.
Use exact enum strings from the schema. Keep missing non-critical list fields as empty arrays.
Required schema:
{
"analysis_summary": "string",
"key_risk_signals": ["string"],
"recommendation": "cautious_reporting | collect_more_evidence | do_not_make_causal_claim",
"recommendation_rationale": ["string"],
"additional_sensitivity_checks": ["string"],
"communication_warnings": ["string"],
"confidence": "low | medium | high"
}
Parser error:
{error_message}
Previous answer:
{raw_output}
'''
).strip()
def compact_report_for_model_comparison(report):
contract = report['analysis_contract']
return {
'estimand': contract['estimand'],
'treatment': contract['treatment'],
'outcome': contract['outcome'],
'placebo_outcome': contract['placebo_outcome'],
'main_estimate': report['observed_adjusted_estimate'],
'oracle_estimate_synthetic_only': report['oracle_hidden_confounder_estimate'],
'placebo_summary': report['placebo_summary'],
'tipping_point': report['tipping_point'],
'sensitivity_grid_summary': report['sensitivity_grid_summary'],
'top_benchmark_shifts': report['benchmark_covariate_table'][:3],
'sensitivity_gates': report['sensitivity_gates'],
'recommended_posture_from_rules': report.get('sensitivity_recommendation', report.get('recommendation', '')),
'brittleness_note': 'Local LLM outputs may vary across reruns; assess schema validity separately from causal reasoning quality.',
}
def build_decision_prompt(report):
schema_hint = {
'analysis_summary': 'string',
'key_risk_signals': ['string'],
'recommendation': 'cautious_reporting | collect_more_evidence | do_not_make_causal_claim',
'recommendation_rationale': ['string'],
'additional_sensitivity_checks': ['string'],
'communication_warnings': ['string'],
'confidence': 'low | medium | high',
}
return textwrap.dedent(
f'''
Produce one compact SensitivityDecision JSON object using this schema.
Use the exact enum strings shown in the schema. Every field is required; use [] when no useful item is available.
Return one JSON object only. Keep each list to 2-4 concise strings.
{json.dumps(schema_hint, indent=2)}
Compact sensitivity report:
{json.dumps(compact_report_for_model_comparison(report), indent=2)}
'''
).strip()
def parse_sensitivity_decision(raw_output):
result = parse_pydantic_output(
raw_output,
SensitivityDecision,
scalar_fields=DECISION_SCALAR_FIELDS,
list_fields=DECISION_LIST_FIELDS,
field_aliases=DECISION_ALIASES,
value_aliases=REVIEW_VALUE_ALIASES,
defaults=DECISION_DEFAULTS,
)
return result.parsed, result.json_text, result.notes
def classify_structured_output_failure(error):
text = clean_generated_text(repr(error)).lower()
if 'empty model output' in text:
return 'empty_output'
if 'field required' in text or 'missing' in text:
return 'missing_required_field'
if 'input should be' in text or 'validation error' in text:
return 'wrong_field_type_or_schema'
if 'invalid json' in text or 'expecting value' in text or 'eof' in text or 'jsondecodeerror' in text:
return 'invalid_json_or_truncated_output'
return 'other_structured_output_error'
def empty_sensitivity_decision():
return SensitivityDecision(
analysis_summary='',
key_risk_signals=[],
recommendation='cautious_reporting',
recommendation_rationale=[],
additional_sensitivity_checks=[],
communication_warnings=[],
confidence='low',
)
def parse_or_repair_decision(raw_output, model_id):
if not clean_generated_text(raw_output):
raise ValueError('empty model output')
try:
parsed, parsed_json, notes = parse_sensitivity_decision(raw_output)
return {
'parsed': parsed,
'parsed_json': parsed_json,
'parser_notes': notes,
'repair_used': bool(notes),
'repair_stage': 'parser' if notes else 'none',
'repaired_raw_output': '',
}
except Exception as first_error:
if not (RUN_SCHEMA_REPAIR_RETRY and RUN_LIVE_LOCAL_LLM):
raise
repair_prompt = DECISION_REPAIR_PROMPT_TEMPLATE.format(
raw_output=clean_generated_text(raw_output)[:5000],
error_message=clean_generated_text(repr(first_error))[:1200],
)
repaired_raw_output = local_chat(
repair_prompt,
system_message=SYSTEM_REVIEW_MESSAGE,
model_id=model_id,
max_new_tokens=900,
temperature=TEMPERATURE,
)
parsed, parsed_json, notes = parse_sensitivity_decision(repaired_raw_output)
return {
'parsed': parsed,
'parsed_json': parsed_json,
'parser_notes': [f'first_parse_error: {classify_structured_output_failure(first_error)}'] + notes,
'repair_used': True,
'repair_stage': 'model_retry',
'repaired_raw_output': repaired_raw_output,
}
def run_single_model_sensitivity_case(label, model_id, role, case):
prompt = build_decision_prompt(case['report'])
raw_output = ''
max_score = score_sensitivity_review(empty_sensitivity_decision(), case['expected_recommendation'])[2]
try:
raw_output = local_chat(
prompt,
system_message=SYSTEM_REVIEW_MESSAGE,
model_id=model_id,
max_new_tokens=900,
temperature=TEMPERATURE,
)
parsed_result = parse_or_repair_decision(raw_output, model_id)
audit, score, max_score = score_sensitivity_review(parsed_result['parsed'], case['expected_recommendation'])
return {
'label': label,
'model_id': model_id,
'role': role,
'case_id': case['case_id'],
'expected_recommendation': case['expected_recommendation'],
'model_recommendation': parsed_result['parsed'].recommendation,
'status': 'ok',
'schema_valid': True,
'repair_used': parsed_result['repair_used'],
'repair_stage': parsed_result['repair_stage'],
'error_type': '',
'review_score': score,
'max_review_score': max_score,
'review_score_share': score / max_score if max_score else 0.0,
'error': '',
'raw_output_preview': clean_generated_text(raw_output)[:500],
}
except Exception as error:
return {
'label': label,
'model_id': model_id,
'role': role,
'case_id': case['case_id'],
'expected_recommendation': case['expected_recommendation'],
'model_recommendation': '',
'status': 'failed',
'schema_valid': False,
'repair_used': False,
'repair_stage': 'failed',
'error_type': classify_structured_output_failure(error),
'review_score': 0,
'max_review_score': max_score,
'review_score_share': 0.0,
'error': clean_generated_text(repr(error))[:900],
'raw_output_preview': clean_generated_text(raw_output)[:500],
}
def summarize_model_results(case_results):
if case_results.empty:
return pd.DataFrame(columns=SUMMARY_COLUMNS)
summary = (
case_results
.groupby(['label', 'model_id', 'role'], as_index=False)
.agg(
cases=('case_id', 'count'),
schema_valid_cases=('schema_valid', 'sum'),
schema_repaired_cases=('repair_used', 'sum'),
mean_review_score=('review_score_share', 'mean'),
failure_types=('error_type', lambda values: sorted({value for value in values if value})),
)
)
summary['schema_reliability'] = summary['schema_valid_cases'] / summary['cases']
return summary[SUMMARY_COLUMNS].sort_values(['mean_review_score', 'schema_reliability'], ascending=False)
def run_all_model_sensitivity_comparison(models_to_compare=MODELS_TO_COMPARE, cases=SENSITIVITY_EVAL_CASES):
rows = []
for label, model_id, role in models_to_compare:
print(f'Running {label}: {model_id}')
for case in cases[:MODEL_COMPARISON_CASE_LIMIT]:
rows.append(run_single_model_sensitivity_case(label, model_id, role, case))
clear_loaded_model_cache()
case_results = pd.DataFrame(rows, columns=CASE_RESULT_COLUMNS)
summary = summarize_model_results(case_results)
return summary, case_results
if RUN_FULL_MODEL_COMPARISON and RUN_LIVE_LOCAL_LLM:
sensitivity_model_summary, sensitivity_case_results = run_all_model_sensitivity_comparison()
else:
sensitivity_model_summary = pd.DataFrame(columns=SUMMARY_COLUMNS)
sensitivity_case_results = pd.DataFrame(columns=CASE_RESULT_COLUMNS)
print('Full model comparison skipped. Set RUN_FULL_MODEL_COMPARISON and RUN_LIVE_LOCAL_LLM to True to run it.')
sensitivity_model_summaryRunning Qwen 0.5B: Qwen/Qwen2.5-0.5B-Instruct
Running Qwen 7B: Qwen/Qwen2.5-7B-Instruct
Running Qwen 14B: Qwen/Qwen2.5-14B-Instruct
Running Qwen 32B: Qwen/Qwen2.5-32B-Instruct
Running Phi mini: microsoft/Phi-3.5-mini-instructRunning Mistral 7B: mistralai/Mistral-7B-Instruct-v0.3
Running Mistral Small 24B: mistralai/Mistral-Small-3.1-24B-Instruct-2503Running Gemma 3 27B: google/gemma-3-27b-it
Running Llama 3.1 8B: meta-llama/Meta-Llama-3.1-8B-Instruct| label | model_id | role | cases | schema_valid_cases | schema_repaired_cases | schema_reliability | mean_review_score | failure_types | |
|---|---|---|---|---|---|---|---|---|---|
| 1 | Llama 3.1 8B | meta-llama/Meta-Llama-3.1-8B-Instruct | industry-standard instruct baseline | 2 | 2 | 0 | 1.0 | 0.95 | [] |
| 0 | Gemma 3 27B | google/gemma-3-27b-it | large non-Qwen comparison | 2 | 2 | 2 | 1.0 | 0.90 | [] |
| 4 | Phi mini | microsoft/Phi-3.5-mini-instruct | compact non-Qwen comparison | 2 | 2 | 2 | 1.0 | 0.90 | [] |
| 3 | Mistral Small 24B | mistralai/Mistral-Small-3.1-24B-Instruct-2503 | strong non-Qwen comparison | 2 | 2 | 2 | 1.0 | 0.85 | [] |
| 7 | Qwen 32B | Qwen/Qwen2.5-32B-Instruct | scale comparison | 2 | 2 | 0 | 1.0 | 0.85 | [] |
| 6 | Qwen 14B | Qwen/Qwen2.5-14B-Instruct | strong local analysis | 2 | 2 | 0 | 1.0 | 0.80 | [] |
| 8 | Qwen 7B | Qwen/Qwen2.5-7B-Instruct | fast default | 2 | 2 | 0 | 1.0 | 0.80 | [] |
| 2 | Mistral 7B | mistralai/Mistral-7B-Instruct-v0.3 | 7B model-family comparison | 2 | 2 | 1 | 1.0 | 0.70 | [] |
| 5 | Qwen 0.5B | Qwen/Qwen2.5-0.5B-Instruct | pipeline smoke test | 2 | 2 | 2 | 1.0 | 0.40 | [] |
Inspecting Case-Level Sensitivity Recommendations
This table shows whether the model recommendation was at least as cautious as the sensitivity gates.
sensitivity_case_results[[
'label',
'case_id',
'expected_recommendation',
'model_recommendation',
'review_score_share',
'status',
'error_type',
]].sort_values(['label', 'case_id']).head(30)| label | case_id | expected_recommendation | model_recommendation | review_score_share | status | error_type | |
|---|---|---|---|---|---|---|---|
| 14 | Gemma 3 27B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.9 | ok | |
| 15 | Gemma 3 27B | strong_hidden_confounding | collect_more_evidence | cautious_reporting | 0.9 | ok | |
| 16 | Llama 3.1 8B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 1.0 | ok | |
| 17 | Llama 3.1 8B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.9 | ok | |
| 10 | Mistral 7B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.6 | ok | |
| 11 | Mistral 7B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.8 | ok | |
| 12 | Mistral Small 24B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.9 | ok | |
| 13 | Mistral Small 24B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.8 | ok | |
| 8 | Phi mini | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.9 | ok | |
| 9 | Phi mini | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.9 | ok | |
| 0 | Qwen 0.5B | moderate_hidden_confounding | cautious_reporting | do_not_make_causal_claim | 0.3 | ok | |
| 1 | Qwen 0.5B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.5 | ok | |
| 4 | Qwen 14B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.8 | ok | |
| 5 | Qwen 14B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.8 | ok | |
| 6 | Qwen 32B | moderate_hidden_confounding | cautious_reporting | cautious_reporting | 0.9 | ok | |
| 7 | Qwen 32B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.8 | ok | |
| 2 | Qwen 7B | moderate_hidden_confounding | cautious_reporting | collect_more_evidence | 0.8 | ok | |
| 3 | Qwen 7B | strong_hidden_confounding | collect_more_evidence | collect_more_evidence | 0.8 | ok |
Inspecting Failed Model Runs
A failed model run receives zero workflow credit instead of producing a NaN score. In sensitivity analysis, invalid or overconfident summaries can hide the most important design warnings.
failed_model_details = sensitivity_case_results.loc[
~sensitivity_case_results['schema_valid'],
['label', 'case_id', 'status', 'error_type', 'error', 'raw_output_preview'],
].reset_index(drop=True)
failed_model_details| label | case_id | status | error_type | error | raw_output_preview |
|---|
Interpreting Repair Counts
Repairs are not automatically failures. They show how much cleanup was needed before the output became usable. A sensitivity-summary assistant that often needs repair should remain firmly under analyst review.
def summarize_repair_stages(case_results):
if case_results.empty or 'repair_stage' not in case_results.columns:
return pd.DataFrame(columns=['repair_stage', 'cases'])
return (
case_results.assign(repair_stage=case_results['repair_stage'].fillna('none'))
.groupby('repair_stage', as_index=False)
.agg(cases=('case_id', 'count'))
.sort_values('cases', ascending=False)
)
summarize_repair_stages(sensitivity_case_results)| repair_stage | cases | |
|---|---|---|
| 0 | none | 9 |
| 1 | parser | 9 |
16. Reusable Sensitivity Analysis Checklist
Before reporting an observational causal estimate, the analyst should be able to answer these questions.
sensitivity_checklist = pd.DataFrame(
[
{'check': 'Estimand and adjustment set are explicit', 'example': analysis_contract['estimand']},
{'check': 'Main estimate is shown with uncertainty', 'example': 'observed adjusted coefficient and robust CI'},
{'check': 'Observed covariates are used as benchmarks', 'example': 'coefficient movement when each covariate is omitted'},
{'check': 'Placebo or negative-control evidence is reviewed', 'example': analysis_contract['placebo_outcome']},
{'check': 'Tipping point is reported', 'example': 'minimum residual-correlation distance to sign reversal'},
{'check': 'Synthetic oracle variables are clearly labeled when present', 'example': 'executive_attention is teaching-only'},
{'check': 'Communication posture matches sensitivity gates', 'example': sensitivity_recommendation},
{'check': 'AI summary is audited', 'example': 'score_sensitivity_review'},
]
)
sensitivity_checklist| check | example | |
|---|---|---|
| 0 | Estimand and adjustment set are explicit | ATE of onboarding_specialist on net_revenue_90d among eligible accounts |
| 1 | Main estimate is shown with uncertainty | observed adjusted coefficient and robust CI |
| 2 | Observed covariates are used as benchmarks | coefficient movement when each covariate is omitted |
| 3 | Placebo or negative-control evidence is reviewed | prior_period_revenue |
| 4 | Tipping point is reported | minimum residual-correlation distance to sign reversal |
| 5 | Synthetic oracle variables are clearly labeled when present | executive_attention is teaching-only |
| 6 | Communication posture matches sensitivity gates | report_with_caution_and_sensitivity_limitations |
| 7 | AI summary is audited | score_sensitivity_review |
17. Exercises
- Increase
hidden_strengthin the simulator and rerun the sensitivity report. How do the placebo and oracle gaps change? - Change the residual-confounding grid from correlations up to 0.60 to correlations up to 0.30. Does a sign reversal still appear?
- Add a second placebo outcome that is less related to
executive_attention. How should interpretation differ? - Modify the gate logic so a large placebo effect automatically triggers
collect_more_evidence. - Ask one local model to summarize the report without explicit warnings. Does it overclaim robustness?
- Replace the continuous outcome with a binary renewal outcome and discuss which sensitivity methods would need to change.
Key Takeaways
Sensitivity analysis is how we keep causal humility operational.
A strong sensitivity workflow has this shape:
- State the estimand and adjustment set.
- Estimate the main effect with uncertainty.
- Benchmark against observed covariates.
- Run placebo or negative-control checks when available.
- Use a residual-confounding grid or other method-specific sensitivity analysis.
- Translate the evidence into a communication posture.
- Use AI to summarize the evidence, then audit whether it stayed cautious.
The practical rule is simple: do not ask whether the result is robust; ask how fragile it is under specific, named violations of the assumptions.