import itertools
import warnings
warnings.filterwarnings("ignore")
from graphviz import Digraph
from IPython.display import Markdown, display
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats
import statsmodels.api as sm
import statsmodels.formula.api as smf
sns.set_theme(style="whitegrid", context="notebook")
pd.set_option("display.max_columns", 120)
pd.set_option("display.float_format", lambda x: f"{x:,.3f}")
def draw_dag(edges, nodes=None, title=None, bidirected=None):
dot = Digraph(graph_attr={"rankdir": "LR"})
dot.attr("node", shape="box", style="rounded,filled", fillcolor="#f7fbff", color="#6baed6")
if title:
dot.attr(label=title, labelloc="t")
if nodes is None:
nodes = sorted(set([v for edge in edges for v in edge]))
for node in nodes:
dot.node(str(node), str(node))
for parent, child in edges:
dot.edge(str(parent), str(child))
if bidirected:
for left, right in bidirected:
dot.edge(str(left), str(right), dir="both", color="#de2d26", style="dashed")
return dot
def residualize(y, controls):
y = np.asarray(y, dtype=float)
if controls is None or controls.shape[1] == 0:
return y - y.mean()
X = sm.add_constant(np.asarray(controls, dtype=float), has_constant="add")
model = sm.OLS(y, X).fit()
return model.resid
def partial_corr_test(df, x, y, cond=()):
cond = list(cond)
if len(cond) == 0:
r, p = stats.pearsonr(df[x], df[y])
return r, p
rx = residualize(df[x], df[cond].to_numpy())
ry = residualize(df[y], df[cond].to_numpy())
r, p = stats.pearsonr(rx, ry)
return r, p
def format_edge(edge):
return " -- ".join(sorted(edge))08. Causal Discovery, With Caveats
Causal discovery asks whether we can learn parts of a causal graph from data.
That is a tempting goal. In industry, we often have hundreds of behavioral, operational, product, and customer variables. We want to know which variables drive which others, where to intervene, and what should be treated as a confounder, mediator, proxy, or outcome.
But causal discovery is not a substitute for causal design. It is a way to generate, constrain, and stress-test causal hypotheses under assumptions.
This notebook is intentionally cautious. We will implement small discovery ideas from scratch, study what they can recover, and then break them with latent confounding, selection, faithfulness failures, measurement choices, and time-ordering mistakes.
Learning Goals
By the end of this notebook, you should be able to:
- Explain what causal discovery tries to learn from observational data.
- Distinguish a DAG, a skeleton, a v-structure, and a Markov equivalence class.
- Explain the causal Markov, faithfulness, causal sufficiency, and acyclicity assumptions.
- Use conditional independence patterns to reason about chains, forks, and colliders.
- Implement a small PC-style skeleton search for linear Gaussian data.
- Explain why many edge directions are not identifiable from conditional independences alone.
- Demonstrate how latent confounding and selection bias can mislead discovery.
- Compare constraint-based, score-based, functional-model, and time-series discovery ideas.
- Use causal discovery responsibly as part of an industry causal workflow.
1. Setup
We will use pandas, numpy, statsmodels, scipy, seaborn, matplotlib, networkx, and Graphviz.
2. What Causal Discovery Is and Is Not
Causal discovery is the problem of learning causal structure from data, often with help from assumptions such as:
- conditional independences,
- temporal order,
- non-Gaussianity,
- additive noise structure,
- sparsity,
- known interventions,
- acyclicity.
It can be useful for:
- generating DAG hypotheses,
- screening variables for follow-up,
- finding surprising conditional independences,
- comparing candidate causal stories,
- designing better experiments.
It should not be used as:
- a button that turns a data table into truth,
- a replacement for subject-matter knowledge,
- proof that an observational association is causal,
- a way to avoid defining interventions and estimands.
discovery_families = pd.DataFrame(
[
{
"family": "Constraint-based",
"core signal": "Conditional independence tests",
"examples": "PC, FCI",
"main caveat": "Sensitive to testing error, hidden variables, faithfulness, and measurement choices.",
},
{
"family": "Score-based",
"core signal": "Search for graph with best penalized likelihood or score",
"examples": "GES, hill-climbing, Bayesian network search",
"main caveat": "Large search space; score equivalence and local optima can obscure direction.",
},
{
"family": "Functional causal models",
"core signal": "Asymmetry in cause-effect data-generating process",
"examples": "LiNGAM, additive noise models",
"main caveat": "Requires strong assumptions such as non-Gaussianity or additive noise.",
},
{
"family": "Continuous optimization",
"core signal": "Optimize over weighted adjacency matrices with acyclicity constraints",
"examples": "NOTEARS-style methods",
"main caveat": "Still depends on modeling assumptions and tuning choices.",
},
{
"family": "Time-series discovery",
"core signal": "Lagged predictive structure and conditional independence over time",
"examples": "Granger-style models, PCMCI",
"main caveat": "Predictive precedence is not automatically structural causality.",
},
]
)
discovery_families| family | core signal | examples | main caveat | |
|---|---|---|---|---|
| 0 | Constraint-based | Conditional independence tests | PC, FCI | Sensitive to testing error, hidden variables, ... |
| 1 | Score-based | Search for graph with best penalized likelihoo... | GES, hill-climbing, Bayesian network search | Large search space; score equivalence and loca... |
| 2 | Functional causal models | Asymmetry in cause-effect data-generating process | LiNGAM, additive noise models | Requires strong assumptions such as non-Gaussi... |
| 3 | Continuous optimization | Optimize over weighted adjacency matrices with... | NOTEARS-style methods | Still depends on modeling assumptions and tuni... |
| 4 | Time-series discovery | Lagged predictive structure and conditional in... | Granger-style models, PCMCI | Predictive precedence is not automatically str... |
3. The Assumption Stack
Causal discovery methods differ, but many rely on some version of the following assumptions.
assumption_table = pd.DataFrame(
[
{
"assumption": "Causal Markov condition",
"meaning": "Each variable is independent of its non-effects given its direct causes.",
"failure mode": "Dependencies do not match the graph.",
},
{
"assumption": "Faithfulness",
"meaning": "All and only graph-implied independences appear in the distribution.",
"failure mode": "Path cancellations hide real causal paths.",
},
{
"assumption": "Causal sufficiency",
"meaning": "There are no unmeasured common causes among measured variables.",
"failure mode": "Latent confounding creates edges or directions that are not direct causes.",
},
{
"assumption": "Acyclicity",
"meaning": "The causal graph has no directed cycles.",
"failure mode": "Feedback systems violate DAG assumptions.",
},
{
"assumption": "Correct variable definitions",
"meaning": "Variables correspond to meaningful causal quantities at the right time scale.",
"failure mode": "Aggregates, proxies, and post-treatment variables distort structure.",
},
{
"assumption": "Independent samples or modeled dependence",
"meaning": "Rows are independent, or dependence is explicitly represented.",
"failure mode": "Network, panel, or time-series dependence creates misleading tests.",
},
]
)
assumption_table| assumption | meaning | failure mode | |
|---|---|---|---|
| 0 | Causal Markov condition | Each variable is independent of its non-effect... | Dependencies do not match the graph. |
| 1 | Faithfulness | All and only graph-implied independences appea... | Path cancellations hide real causal paths. |
| 2 | Causal sufficiency | There are no unmeasured common causes among me... | Latent confounding creates edges or directions... |
| 3 | Acyclicity | The causal graph has no directed cycles. | Feedback systems violate DAG assumptions. |
| 4 | Correct variable definitions | Variables correspond to meaningful causal quan... | Aggregates, proxies, and post-treatment variab... |
| 5 | Independent samples or modeled dependence | Rows are independent, or dependence is explici... | Network, panel, or time-series dependence crea... |
The key professional habit is to state the assumptions before showing the graph.
A discovered graph should usually be labeled:
Candidate causal structure under assumptions A, B, and C.
4. Chains, Forks, and Colliders
Most graphical discovery logic starts with conditional independence.
Three basic structures matter:
- Chain: \(X \rightarrow Z \rightarrow Y\)
- Fork: \(X \leftarrow Z \rightarrow Y\)
- Collider: \(X \rightarrow Z \leftarrow Y\)
Chains and forks imply:
\[ X \not\!\perp\!\!\!\perp Y \]
but:
\[ X \perp\!\!\!\perp Y \mid Z \]
Colliders imply the opposite pattern:
\[ X \perp\!\!\!\perp Y \]
but:
\[ X \not\!\perp\!\!\!\perp Y \mid Z \]
rng = np.random.default_rng(4808)
n = 5000
chain = pd.DataFrame({"X": rng.normal(size=n)})
chain["Z"] = 0.9 * chain["X"] + rng.normal(size=n)
chain["Y"] = 0.9 * chain["Z"] + rng.normal(size=n)
fork = pd.DataFrame({"Z": rng.normal(size=n)})
fork["X"] = 0.9 * fork["Z"] + rng.normal(size=n)
fork["Y"] = 0.9 * fork["Z"] + rng.normal(size=n)
collider = pd.DataFrame({"X": rng.normal(size=n), "Y": rng.normal(size=n)})
collider["Z"] = 0.9 * collider["X"] + 0.9 * collider["Y"] + rng.normal(size=n)
ci_rows = []
for name, df in [("Chain X -> Z -> Y", chain), ("Fork X <- Z -> Y", fork), ("Collider X -> Z <- Y", collider)]:
marginal_r, marginal_p = partial_corr_test(df, "X", "Y", cond=())
conditional_r, conditional_p = partial_corr_test(df, "X", "Y", cond=("Z",))
ci_rows.extend(
[
{
"structure": name,
"test": "marginal X-Y",
"correlation": marginal_r,
"p_value": marginal_p,
},
{
"structure": name,
"test": "conditional X-Y | Z",
"correlation": conditional_r,
"p_value": conditional_p,
},
]
)
ci_table = pd.DataFrame(ci_rows)
ci_table| structure | test | correlation | p_value | |
|---|---|---|---|---|
| 0 | Chain X -> Z -> Y | marginal X-Y | 0.500 | 0.000 |
| 1 | Chain X -> Z -> Y | conditional X-Y | Z | -0.022 | 0.112 |
| 2 | Fork X <- Z -> Y | marginal X-Y | 0.459 | 0.000 |
| 3 | Fork X <- Z -> Y | conditional X-Y | Z | 0.015 | 0.302 |
| 4 | Collider X -> Z <- Y | marginal X-Y | -0.001 | 0.952 |
| 5 | Collider X -> Z <- Y | conditional X-Y | Z | -0.462 | 0.000 |
graphs = [
draw_dag([("X", "Z"), ("Z", "Y")], title="Chain"),
draw_dag([("Z", "X"), ("Z", "Y")], title="Fork"),
draw_dag([("X", "Z"), ("Y", "Z")], title="Collider"),
]
for graph in graphs:
display(graph)fig, ax = plt.subplots(figsize=(8.5, 4))
plot_df = ci_table.copy()
plot_df["abs_correlation"] = plot_df["correlation"].abs()
sns.barplot(data=plot_df, y="structure", x="abs_correlation", hue="test", ax=ax)
ax.set_title("Conditioning has opposite implications for colliders")
ax.set_xlabel("Absolute correlation")
ax.set_ylabel("")
plt.tight_layout()
plt.show()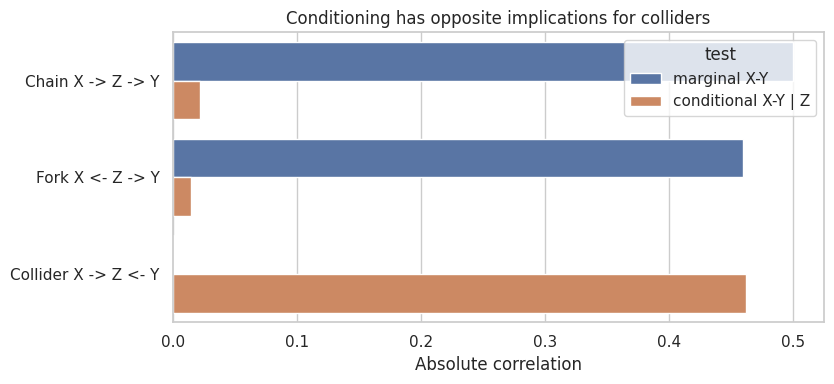
This is the heart of many discovery methods:
- remove edges when conditional independence appears,
- orient unshielded colliders when conditioning behavior requires it,
- leave unresolved directions when multiple DAGs imply the same independences.
5. Markov Equivalence
Different DAGs can imply the same conditional independence structure.
For example:
\[ X \rightarrow Z \rightarrow Y \]
and:
\[ X \leftarrow Z \rightarrow Y \]
both imply \(X \perp\!\!\!\perp Y \mid Z\). Conditional independences alone cannot choose between them.
But:
\[ X \rightarrow Z \leftarrow Y \]
has a collider at \(Z\), so it is distinguishable from the chain and fork.
equivalence_table = pd.DataFrame(
[
{
"DAG": "X -> Z -> Y",
"skeleton": "X - Z - Y",
"unshielded collider?": "No",
"key independence": "X independent of Y given Z",
"equivalence class": "Same as fork",
},
{
"DAG": "X <- Z -> Y",
"skeleton": "X - Z - Y",
"unshielded collider?": "No",
"key independence": "X independent of Y given Z",
"equivalence class": "Same as chain",
},
{
"DAG": "X -> Z <- Y",
"skeleton": "X - Z - Y",
"unshielded collider?": "Yes",
"key independence": "X independent of Y marginally, dependent given Z",
"equivalence class": "Distinct",
},
]
)
equivalence_table| DAG | skeleton | unshielded collider? | key independence | equivalence class | |
|---|---|---|---|---|---|
| 0 | X -> Z -> Y | X - Z - Y | No | X independent of Y given Z | Same as fork |
| 1 | X <- Z -> Y | X - Z - Y | No | X independent of Y given Z | Same as chain |
| 2 | X -> Z <- Y | X - Z - Y | Yes | X independent of Y marginally, dependent given Z | Distinct |
This is why many discovery algorithms output a partially directed graph rather than a fully directed DAG. The data may identify an equivalence class, not one unique causal story.
6. A Small PC-Style Skeleton Search
The PC algorithm is a classic constraint-based causal discovery method. The full algorithm has several steps. Here we implement a tiny version focused on the skeleton:
- Start with a complete undirected graph.
- Test marginal independences and remove edges.
- Test conditional independences with increasingly large conditioning sets.
- Record the separating set that removed each edge.
- Use separating sets to orient unshielded colliders.
This is a teaching implementation, not a production implementation.
def neighbors(edge_set, node):
out = set()
for left, right in edge_set:
if left == node:
out.add(right)
elif right == node:
out.add(left)
return out
def canonical_pair(x, y):
return tuple(sorted((x, y)))
def pc_skeleton(df, variables, alpha=0.01, max_cond_set=3):
edge_set = {canonical_pair(x, y) for x, y in itertools.combinations(variables, 2)}
sep_sets = {}
test_log = []
for cond_size in range(max_cond_set + 1):
removed_this_round = True
while removed_this_round:
removed_this_round = False
for x, y in list(edge_set):
candidate_neighbors = list((neighbors(edge_set, x) - {y}) | (neighbors(edge_set, y) - {x}))
if len(candidate_neighbors) < cond_size:
continue
for cond in itertools.combinations(candidate_neighbors, cond_size):
r, p = partial_corr_test(df, x, y, cond=cond)
test_log.append(
{
"x": x,
"y": y,
"conditioning_set": cond,
"cond_size": cond_size,
"partial_corr": r,
"p_value": p,
}
)
if p > alpha:
edge_set.remove((x, y))
sep_sets[canonical_pair(x, y)] = tuple(cond)
removed_this_round = True
break
return edge_set, sep_sets, pd.DataFrame(test_log)
def orient_v_structures(variables, edge_set, sep_sets):
directed = []
undirected = set(edge_set)
for x, z in itertools.combinations(variables, 2):
if canonical_pair(x, z) in edge_set:
continue
common = neighbors(edge_set, x) & neighbors(edge_set, z)
sep = set(sep_sets.get(canonical_pair(x, z), ()))
for y in common:
if y not in sep:
directed.append((x, y))
directed.append((z, y))
undirected.discard(canonical_pair(x, y))
undirected.discard(canonical_pair(z, y))
return directed, undirected
def draw_partial_graph(variables, directed_edges, undirected_edges, title=None):
dot = Digraph(graph_attr={"rankdir": "LR"})
dot.attr("node", shape="box", style="rounded,filled", fillcolor="#f7fbff", color="#6baed6")
if title:
dot.attr(label=title, labelloc="t")
for node in variables:
dot.node(node, node)
for left, right in sorted(undirected_edges):
dot.edge(left, right, dir="none", color="#636363")
for left, right in directed_edges:
dot.edge(left, right, color="#2b8cbe")
return dotNow simulate a linear Gaussian DAG:
\[ A \rightarrow B \]
\[ A \rightarrow C \]
\[ B \rightarrow D \]
\[ C \rightarrow D \]
\[ D \rightarrow E \]
rng = np.random.default_rng(4809)
n = 4500
dag_df = pd.DataFrame({"A": rng.normal(size=n)})
dag_df["B"] = 0.90 * dag_df["A"] + rng.normal(size=n)
dag_df["C"] = 0.75 * dag_df["A"] + rng.normal(size=n)
dag_df["D"] = 0.70 * dag_df["B"] + 0.65 * dag_df["C"] + rng.normal(size=n)
dag_df["E"] = 0.85 * dag_df["D"] + rng.normal(size=n)
variables = ["A", "B", "C", "D", "E"]
true_edges = {canonical_pair(*edge) for edge in [("A", "B"), ("A", "C"), ("B", "D"), ("C", "D"), ("D", "E")]}
edge_set, sep_sets, pc_log = pc_skeleton(dag_df, variables, alpha=0.001, max_cond_set=3)
directed_edges, remaining_undirected = orient_v_structures(variables, edge_set, sep_sets)
comparison = pd.DataFrame(
[
{
"edge": format_edge(edge),
"true_edge": edge in true_edges,
"pc_skeleton_edge": edge in edge_set,
}
for edge in sorted(set(true_edges) | set(edge_set))
]
)
draw_dag([("A", "B"), ("A", "C"), ("B", "D"), ("C", "D"), ("D", "E")], title="True data-generating DAG")display(draw_partial_graph(variables, directed_edges, remaining_undirected, title="Recovered partial graph"))
comparison| edge | true_edge | pc_skeleton_edge | |
|---|---|---|---|
| 0 | A -- B | True | True |
| 1 | A -- C | True | True |
| 2 | B -- D | True | True |
| 3 | C -- D | True | True |
| 4 | D -- E | True | True |
sep_table = pd.DataFrame(
[
{
"removed_pair": f"{x} -- {y}",
"separating_set": ", ".join(cond) if cond else "(empty)",
}
for (x, y), cond in sep_sets.items()
]
).sort_values("removed_pair")
sep_table| removed_pair | separating_set | |
|---|---|---|
| 4 | A -- D | C, B |
| 2 | A -- E | D |
| 0 | B -- C | A |
| 3 | B -- E | D |
| 1 | C -- E | D |
Interpretation
In this friendly simulation, the skeleton search recovers the main adjacencies. It also orients the unshielded collider:
\[ B \rightarrow D \leftarrow C \]
because \(B\) and \(C\) are not adjacent and \(D\) was not in their separating set.
But several directions remain unresolved. This is not a failure of the implementation. Conditional independence alone often identifies a Markov equivalence class, not a unique DAG.
7. Testing Error and Sample Size
Conditional independence testing is statistical. With finite data:
- true edges can be removed,
- false edges can remain,
- orientation decisions can change,
- results can be sensitive to the significance threshold.
Let us rerun the same skeleton procedure with smaller sample sizes.
def simulate_friendly_dag(n, seed):
rng = np.random.default_rng(seed)
out = pd.DataFrame({"A": rng.normal(size=n)})
out["B"] = 0.90 * out["A"] + rng.normal(size=n)
out["C"] = 0.75 * out["A"] + rng.normal(size=n)
out["D"] = 0.70 * out["B"] + 0.65 * out["C"] + rng.normal(size=n)
out["E"] = 0.85 * out["D"] + rng.normal(size=n)
return out
sample_rows = []
for sample_size in [150, 300, 800, 2000, 4500]:
for rep in range(20):
tmp = simulate_friendly_dag(sample_size, seed=10_000 + sample_size + rep)
edges_hat, _, _ = pc_skeleton(tmp, variables, alpha=0.001, max_cond_set=3)
true_positive = len(edges_hat & true_edges)
false_positive = len(edges_hat - true_edges)
false_negative = len(true_edges - edges_hat)
sample_rows.append(
{
"sample_size": sample_size,
"rep": rep,
"true_positive_edges": true_positive,
"false_positive_edges": false_positive,
"false_negative_edges": false_negative,
}
)
sample_sensitivity = pd.DataFrame(sample_rows)
sample_summary = sample_sensitivity.groupby("sample_size", as_index=False).agg(
mean_true_positive=("true_positive_edges", "mean"),
mean_false_positive=("false_positive_edges", "mean"),
mean_false_negative=("false_negative_edges", "mean"),
)
sample_summary| sample_size | mean_true_positive | mean_false_positive | mean_false_negative | |
|---|---|---|---|---|
| 0 | 150 | 4.350 | 0.000 | 0.650 |
| 1 | 300 | 4.950 | 0.000 | 0.050 |
| 2 | 800 | 5.000 | 0.000 | 0.000 |
| 3 | 2000 | 5.000 | 0.000 | 0.000 |
| 4 | 4500 | 5.000 | 0.000 | 0.000 |
plot_df = sample_summary.melt(
id_vars="sample_size",
value_vars=["mean_true_positive", "mean_false_positive", "mean_false_negative"],
var_name="metric",
value_name="mean_count",
)
fig, ax = plt.subplots(figsize=(8, 4.2))
sns.lineplot(data=plot_df, x="sample_size", y="mean_count", hue="metric", marker="o", ax=ax)
ax.set_title("Discovery quality changes with sample size")
ax.set_xlabel("Sample size")
ax.set_ylabel("Mean edge count across repetitions")
plt.tight_layout()
plt.show()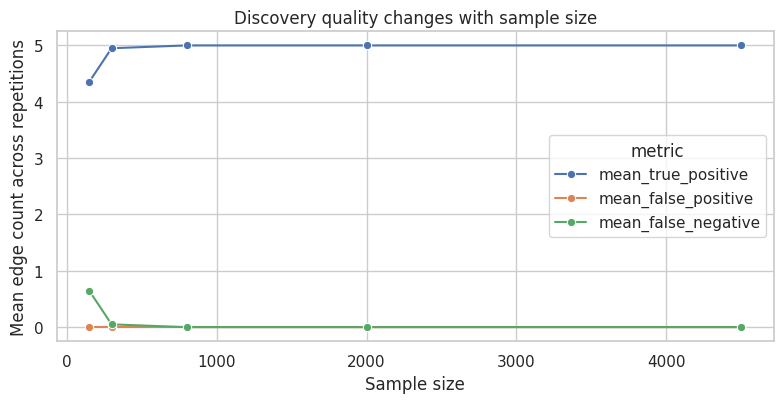
This is one reason discovery should be paired with stability checks. If small changes in sample, threshold, or variable set produce very different graphs, the discovered structure should be treated as exploratory.
8. Score-Based Search: A Tiny Exhaustive Example
Score-based methods search over graphs and choose a graph with a good penalized fit.
For a small number of variables, we can enumerate all possible directed graphs, keep only acyclic ones, and score each graph with a linear-Gaussian BIC:
\[ BIC = \sum_j \left[ n \log\left(\frac{RSS_j}{n}\right) + k_j \log(n) \right] \]
where \(RSS_j\) is the residual sum of squares for node \(j\) regressed on its parents.
score_vars = ["A", "B", "C", "D"]
score_df = dag_df[score_vars].copy()
def graph_bic(df, directed_edges, variables):
n = len(df)
total = 0.0
for child in variables:
parents = [p for p, c in directed_edges if c == child]
y = df[child].to_numpy()
if parents:
X = sm.add_constant(df[parents].to_numpy(), has_constant="add")
else:
X = np.ones((n, 1))
model = sm.OLS(y, X).fit()
rss = np.sum(model.resid ** 2)
k = X.shape[1]
total += n * np.log(rss / n) + k * np.log(n)
return total
def enumerate_dags(df, variables, max_parents=3):
possible_edges = [(a, b) for a in variables for b in variables if a != b]
rows = []
for mask in range(2 ** len(possible_edges)):
edges = [possible_edges[i] for i in range(len(possible_edges)) if (mask >> i) & 1]
parent_counts = pd.Series([child for _, child in edges]).value_counts()
if len(parent_counts) and parent_counts.max() > max_parents:
continue
G = nx.DiGraph()
G.add_nodes_from(variables)
G.add_edges_from(edges)
if not nx.is_directed_acyclic_graph(G):
continue
rows.append(
{
"edges": tuple(sorted(edges)),
"n_edges": len(edges),
"bic": graph_bic(df, edges, variables),
}
)
return pd.DataFrame(rows).sort_values("bic").reset_index(drop=True)
score_results = enumerate_dags(score_df, score_vars, max_parents=3)
top_score_graphs = score_results.head(8).copy()
top_score_graphs["edge_text"] = top_score_graphs["edges"].apply(lambda edges: ", ".join([f"{a}->{b}" for a, b in edges]))
top_score_graphs[["bic", "n_edges", "edge_text"]]| bic | n_edges | edge_text | |
|---|---|---|---|
| 0 | -175.489 | 4 | A->B, A->C, B->D, C->D |
| 1 | -175.489 | 4 | A->B, B->D, C->A, C->D |
| 2 | -175.489 | 4 | A->C, B->A, B->D, C->D |
| 3 | -168.405 | 5 | A->B, A->C, A->D, B->D, C->D |
| 4 | -168.405 | 5 | A->B, A->D, B->D, C->A, C->D |
| 5 | -168.405 | 5 | A->C, A->D, B->A, B->D, C->D |
| 6 | -168.365 | 5 | A->B, A->C, B->C, B->D, C->D |
| 7 | -168.365 | 5 | B->A, B->C, B->D, C->A, D->C |
best_edges = list(score_results.iloc[0]["edges"])
display(draw_dag(best_edges, nodes=score_vars, title="Best BIC DAG among enumerated candidates"))
pd.Series(
{
"true_edges_on_A_B_C_D": "A->B, A->C, B->D, C->D",
"best_score_edges": ", ".join([f"{a}->{b}" for a, b in best_edges]),
"number_of_acyclic_candidates_scored": len(score_results),
}
)true_edges_on_A_B_C_D A->B, A->C, B->D, C->D
best_score_edges A->B, A->C, B->D, C->D
number_of_acyclic_candidates_scored 543
dtype: objectThe score-based search can recover a plausible graph in this friendly setting. But score-based search has its own caveats:
- many DAGs can score similarly,
- the best score depends on modeling assumptions,
- the search space grows rapidly,
- equivalent structures can be hard to distinguish,
- omitted variables can make a well-scored graph causally wrong.
9. Functional Causal Models: Direction From Asymmetry
Some methods use stronger assumptions to orient edges that conditional independence cannot orient.
LiNGAM assumes a linear acyclic model with independent non-Gaussian errors and no unobserved confounding. Under those assumptions, the direction can be identifiable.
Here is a simple two-variable illustration. We simulate:
\[ X \rightarrow Y \]
with non-Gaussian \(X\) and independent noise:
\[ Y = 1.7X + \epsilon \]
If the model is correct, the residual from \(Y\) regressed on \(X\) should be independent of \(X\). The reverse regression usually leaves a residual that still depends on \(Y\).
def distance_correlation(x, y):
x = np.asarray(x, dtype=float).reshape(-1, 1)
y = np.asarray(y, dtype=float).reshape(-1, 1)
a = np.abs(x - x.T)
b = np.abs(y - y.T)
A = a - a.mean(axis=0, keepdims=True) - a.mean(axis=1, keepdims=True) + a.mean()
B = b - b.mean(axis=0, keepdims=True) - b.mean(axis=1, keepdims=True) + b.mean()
dcov2 = np.mean(A * B)
dvarx = np.mean(A * A)
dvary = np.mean(B * B)
return np.sqrt(max(dcov2, 0) / np.sqrt(dvarx * dvary))
rng = np.random.default_rng(4810)
n = 1200
X = rng.laplace(loc=0, scale=1, size=n)
Y = 1.7 * X + rng.normal(0, 1.0, size=n)
fm_df = pd.DataFrame({"X": X, "Y": Y})
forward = smf.ols("Y ~ X", data=fm_df).fit()
reverse = smf.ols("X ~ Y", data=fm_df).fit()
dependence_summary = pd.DataFrame(
[
{
"direction": "Assume X -> Y",
"residual": "Y residual after regressing on X",
"dependence_between_predictor_and_residual": distance_correlation(fm_df["X"], forward.resid),
},
{
"direction": "Assume Y -> X",
"residual": "X residual after regressing on Y",
"dependence_between_predictor_and_residual": distance_correlation(fm_df["Y"], reverse.resid),
},
]
)
dependence_summary| direction | residual | dependence_between_predictor_and_residual | |
|---|---|---|---|
| 0 | Assume X -> Y | Y residual after regressing on X | 0.045 |
| 1 | Assume Y -> X | X residual after regressing on Y | 0.095 |
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
sns.scatterplot(x=fm_df["X"], y=forward.resid, alpha=0.35, ax=axes[0])
axes[0].set_title("Forward residuals are closer to independent")
axes[0].set_xlabel("X")
axes[0].set_ylabel("Residual from Y ~ X")
sns.scatterplot(x=fm_df["Y"], y=reverse.resid, alpha=0.35, ax=axes[1])
axes[1].set_title("Reverse residuals retain dependence")
axes[1].set_xlabel("Y")
axes[1].set_ylabel("Residual from X ~ Y")
plt.tight_layout()
plt.show()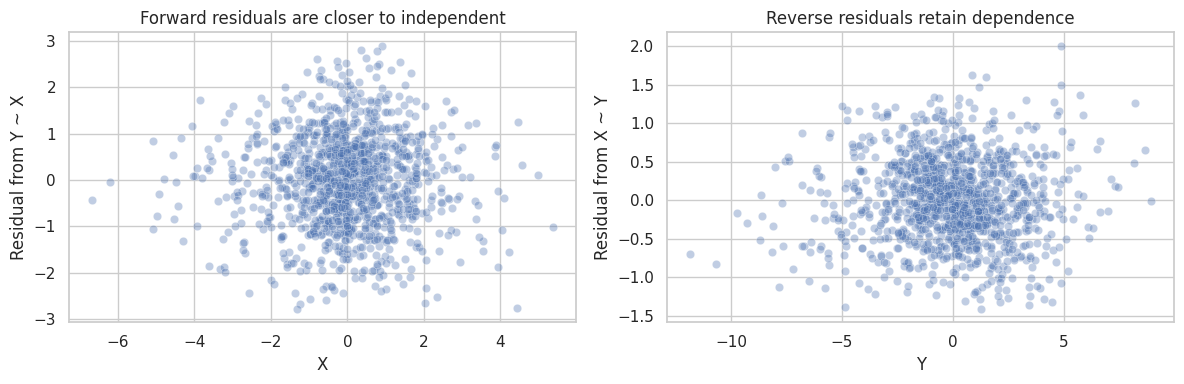
This kind of asymmetry is useful, but only under strong conditions. If the relationship is nonlinear, the noise is not independent, variables are coarsened, or there is an unmeasured common cause, the diagnostic can mislead.
10. Failure Mode: Latent Confounding
Now introduce an unmeasured variable \(U\):
\[ U \rightarrow X \]
and:
\[ U \rightarrow Y \]
If \(U\) is not in the dataset, observational data can make \(X\) and \(Y\) look directly connected. A discovery algorithm may include an edge between them, but the edge may represent unmeasured confounding rather than a direct causal effect.
rng = np.random.default_rng(4811)
n = 4000
latent = pd.DataFrame({"U": rng.normal(size=n)})
latent["X"] = 1.1 * latent["U"] + rng.normal(size=n)
latent["Y"] = 1.2 * latent["U"] + rng.normal(size=n)
latent["Z"] = 0.9 * latent["Y"] + rng.normal(size=n)
observed_latent = latent[["X", "Y", "Z"]]
latent_edges, latent_sep, _ = pc_skeleton(observed_latent, ["X", "Y", "Z"], alpha=0.001, max_cond_set=2)
latent_directed, latent_undirected = orient_v_structures(["X", "Y", "Z"], latent_edges, latent_sep)
display(draw_dag([("U", "X"), ("U", "Y"), ("Y", "Z")], nodes=["U", "X", "Y", "Z"], title="True graph includes unobserved U"))
display(draw_partial_graph(["X", "Y", "Z"], latent_directed, latent_undirected, title="Observed-variable discovery result"))
pd.DataFrame(
[
{
"observed_pair": format_edge(edge),
"edge_found": True,
"causal_warning": "Could be direct causation, latent confounding, or both.",
}
for edge in sorted(latent_edges)
]
)| observed_pair | edge_found | causal_warning | |
|---|---|---|---|
| 0 | X -- Y | True | Could be direct causation, latent confounding,... |
| 1 | Y -- Z | True | Could be direct causation, latent confounding,... |
FCI-style algorithms are designed to represent possible latent confounding more explicitly than PC. But no algorithm can recover information that the data and assumptions do not contain.
11. Failure Mode: Selection Bias
Conditioning on a collider can create associations.
Suppose:
\[ X \rightarrow S \leftarrow Y \]
where \(S\) means “included in the dataset.” If we only observe rows with \(S=1\), then \(X\) and \(Y\) can become associated even if neither causes the other.
This is common in industry:
- only converted users enter a downstream dataset,
- only approved loans are observed,
- only active customers have telemetry,
- only investigated transactions receive fraud labels.
rng = np.random.default_rng(4812)
n = 80_000
selection = pd.DataFrame({"X": rng.normal(size=n), "Y": rng.normal(size=n)})
selection["selection_score"] = 0.9 * selection["X"] + 0.9 * selection["Y"] + rng.normal(size=n)
threshold = selection["selection_score"].quantile(0.82)
selection["S"] = (selection["selection_score"] > threshold).astype(int)
selected = selection.loc[selection["S"] == 1].copy()
selection_summary = pd.DataFrame(
[
{
"sample": "Full generated population",
"n": len(selection),
"corr_XY": selection[["X", "Y"]].corr().iloc[0, 1],
},
{
"sample": "Selected sample S=1",
"n": len(selected),
"corr_XY": selected[["X", "Y"]].corr().iloc[0, 1],
},
]
)
selection_summary| sample | n | corr_XY | |
|---|---|---|---|
| 0 | Full generated population | 80000 | -0.006 |
| 1 | Selected sample S=1 | 14400 | -0.331 |
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
sns.scatterplot(data=selection.sample(2500, random_state=1), x="X", y="Y", alpha=0.25, ax=axes[0])
axes[0].set_title("Full population: X and Y are independent")
sns.scatterplot(data=selected.sample(2500, random_state=2), x="X", y="Y", alpha=0.25, ax=axes[1], color="#2b8cbe")
axes[1].set_title("Selected sample: collider conditioning creates association")
plt.tight_layout()
plt.show()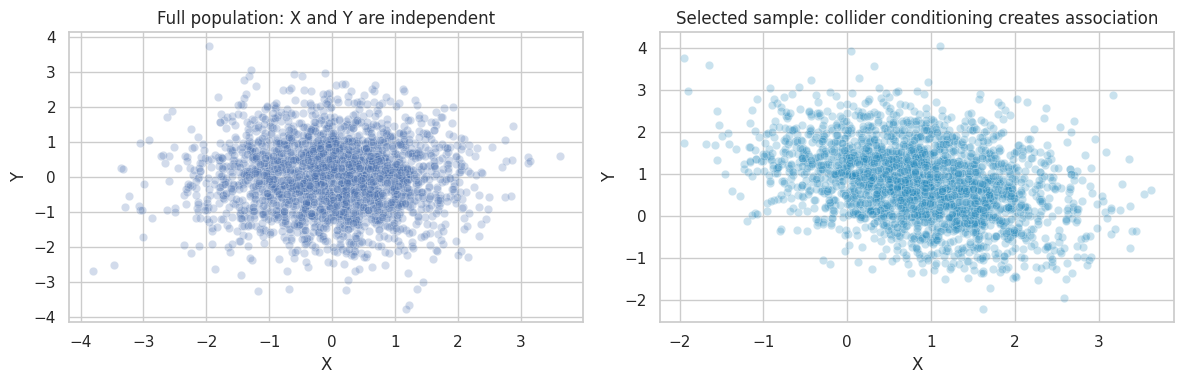
If causal discovery is run only on the selected sample, it can interpret selection-induced dependence as structure. Before discovery, define the data-generating pipeline, not only the variables.
12. Failure Mode: Faithfulness Cancellation
Faithfulness can fail when multiple causal paths cancel.
Suppose:
\[ X \rightarrow Y \]
and:
\[ X \rightarrow M \rightarrow Y \]
If the direct path is positive and the indirect path is negative with similar magnitude, the marginal association between \(X\) and \(Y\) can be near zero even though causal effects exist.
rng = np.random.default_rng(4813)
n = 5000
faith = pd.DataFrame({"X": rng.normal(size=n)})
faith["M"] = 1.0 * faith["X"] + rng.normal(0, 0.3, size=n)
faith["Y"] = 1.0 * faith["X"] - 1.0 * faith["M"] + rng.normal(0, 0.3, size=n)
faith_tests = pd.DataFrame(
[
{
"test": "Marginal X-Y",
"correlation": partial_corr_test(faith, "X", "Y")[0],
"p_value": partial_corr_test(faith, "X", "Y")[1],
},
{
"test": "Conditional X-Y | M",
"correlation": partial_corr_test(faith, "X", "Y", cond=("M",))[0],
"p_value": partial_corr_test(faith, "X", "Y", cond=("M",))[1],
},
{
"test": "Marginal X-M",
"correlation": partial_corr_test(faith, "X", "M")[0],
"p_value": partial_corr_test(faith, "X", "M")[1],
},
{
"test": "Marginal M-Y",
"correlation": partial_corr_test(faith, "M", "Y")[0],
"p_value": partial_corr_test(faith, "M", "Y")[1],
},
]
)
display(draw_dag([("X", "Y"), ("X", "M"), ("M", "Y")], title="Faithfulness can fail by cancellation"))
faith_tests| test | correlation | p_value | |
|---|---|---|---|
| 0 | Marginal X-Y | 0.022 | 0.122 |
| 1 | Conditional X-Y | M | 0.695 | 0.000 |
| 2 | Marginal X-M | 0.958 | 0.000 |
| 3 | Marginal M-Y | -0.183 | 0.000 |
The graph contains a direct causal path from \(X\) to \(Y\), but the marginal correlation is close to zero. Algorithms that rely on observed independences can remove real causal edges when faithfulness fails.
Exact cancellation is special, but near-cancellation is not exotic in business systems with offsetting mechanisms.
13. Time Series: Predictive Precedence Is Not Enough
Time order helps, but it does not solve everything.
If \(X_t\) predicts \(Y_{t+1}\), that is useful. But it may reflect:
- \(X_t \rightarrow Y_{t+1}\),
- a common driver,
- seasonality,
- feedback,
- measurement timing,
- autocorrelation.
Time-series causal discovery needs assumptions about lags, stationarity, sampling frequency, hidden variables, and feedback.
rng = np.random.default_rng(4814)
T = 500
ts_rows = []
u_prev = 0
x_prev = 0
y_prev = 0
for t in range(T):
U = 0.7 * u_prev + rng.normal()
X = 0.5 * x_prev + 0.9 * U + rng.normal()
Y = 0.5 * y_prev + 0.8 * U + 0.0 * x_prev + rng.normal()
ts_rows.append({"time": t, "U": U, "X": X, "Y": Y})
u_prev, x_prev, y_prev = U, X, Y
ts = pd.DataFrame(ts_rows)
ts["X_lag"] = ts["X"].shift(1)
ts["Y_lag"] = ts["Y"].shift(1)
ts["U_lag"] = ts["U"].shift(1)
ts_model_naive = smf.ols("Y ~ Y_lag + X_lag", data=ts.dropna()).fit()
ts_model_with_driver = smf.ols("Y ~ Y_lag + X_lag + U_lag", data=ts.dropna()).fit()
ts_results = pd.DataFrame(
{
"Lag model without common driver": {
"estimate": ts_model_naive.params["X_lag"],
"std_error": ts_model_naive.bse["X_lag"],
"p_value": ts_model_naive.pvalues["X_lag"],
},
"Lag model with common driver observed": {
"estimate": ts_model_with_driver.params["X_lag"],
"std_error": ts_model_with_driver.bse["X_lag"],
"p_value": ts_model_with_driver.pvalues["X_lag"],
},
}
).T
ts_results| estimate | std_error | p_value | |
|---|---|---|---|
| Lag model without common driver | 0.167 | 0.038 | 0.000 |
| Lag model with common driver observed | 0.011 | 0.044 | 0.801 |
fig, ax = plt.subplots(figsize=(8.5, 4))
plot_ts = ts.iloc[:140].copy()
sns.lineplot(data=plot_ts, x="time", y="X", ax=ax, label="X")
sns.lineplot(data=plot_ts, x="time", y="Y", ax=ax, label="Y")
sns.lineplot(data=plot_ts, x="time", y="U", ax=ax, label="Common driver U", alpha=0.65)
ax.set_title("Lagged prediction can reflect a common time-varying driver")
ax.set_xlabel("Time")
ax.set_ylabel("Value")
plt.tight_layout()
plt.show()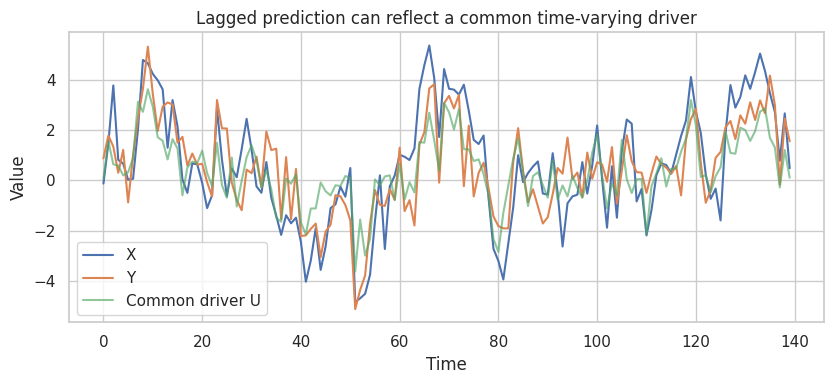
The naive lag model can make \(X\) look predictive of future \(Y\) even when the true structural coefficient from lagged \(X\) to \(Y\) is zero. Once the common driver is included, the apparent lag effect shrinks.
In real work, the common driver may be unobserved. That is why time order is helpful but not definitive.
14. Discovery Plus Interventions
Causal discovery is strongest when paired with interventions.
A practical workflow is:
- Use observational data to propose a small number of plausible graphs.
- Use domain constraints to rule out impossible directions.
- Identify high-value uncertain edges.
- Run an experiment or quasi-experiment to test those edges.
- Update the graph and the intervention policy.
rng = np.random.default_rng(4815)
n = 6000
obs = pd.DataFrame({"Marketing": rng.normal(size=n)})
obs["Traffic"] = 0.9 * obs["Marketing"] + rng.normal(size=n)
obs["Revenue"] = 1.1 * obs["Traffic"] + 0.4 * obs["Marketing"] + rng.normal(size=n)
obs_edges, obs_sep, _ = pc_skeleton(obs, ["Marketing", "Traffic", "Revenue"], alpha=0.001, max_cond_set=2)
obs_directed, obs_undirected = orient_v_structures(["Marketing", "Traffic", "Revenue"], obs_edges, obs_sep)
# Intervention: randomize traffic boost while holding marketing independent of assignment.
experiment = pd.DataFrame({"Marketing": rng.normal(size=n)})
experiment["TrafficBoost"] = rng.binomial(1, 0.5, size=n)
experiment["Traffic"] = 0.9 * experiment["Marketing"] + 1.0 * experiment["TrafficBoost"] + rng.normal(size=n)
experiment["Revenue"] = 1.1 * experiment["Traffic"] + 0.4 * experiment["Marketing"] + rng.normal(size=n)
experiment_model = smf.ols("Revenue ~ TrafficBoost + Marketing", data=experiment).fit()
experiment_summary = pd.Series(
{
"observational_candidate_edges": ", ".join([format_edge(edge) for edge in sorted(obs_edges)]),
"experiment_effect_of_traffic_boost_on_revenue": experiment_model.params["TrafficBoost"],
"experiment_p_value": experiment_model.pvalues["TrafficBoost"],
}
)
display(draw_partial_graph(["Marketing", "Traffic", "Revenue"], obs_directed, obs_undirected, title="Observational candidate graph"))
experiment_summaryobservational_candidate_edges Marketing -- Revenue, Marketing -- Traffic, Re...
experiment_effect_of_traffic_boost_on_revenue 1.073
experiment_p_value 0.000
dtype: objectThe observational graph can suggest a causal story. The intervention tests a key part of that story. This is the productive relationship: discovery helps prioritize experiments; experiments discipline discovery.
15. Industry Workflow for Causal Discovery
A responsible workflow looks more like engineering due diligence than automated graph generation.
workflow = pd.DataFrame(
[
{
"step": "Define the variable dictionary",
"question": "What does each variable mean, when is it measured, and could it be post-treatment?",
"artifact": "Measurement and timing table.",
},
{
"step": "Encode hard constraints",
"question": "Which directions are impossible because of time, product logic, or business rules?",
"artifact": "Allowed/forbidden edge list.",
},
{
"step": "Choose an assumption family",
"question": "Are we relying on conditional independences, scores, non-Gaussianity, time order, or interventions?",
"artifact": "Discovery assumptions memo.",
},
{
"step": "Run stability checks",
"question": "Does the graph change under resampling, thresholds, variable subsets, or alternative algorithms?",
"artifact": "Edge stability table.",
},
{
"step": "Audit caveats",
"question": "Could hidden confounders, selection, measurement error, or feedback explain key edges?",
"artifact": "Threat model.",
},
{
"step": "Prioritize interventions",
"question": "Which uncertain edge would change the decision if confirmed or rejected?",
"artifact": "Experiment backlog.",
},
{
"step": "Report as candidate structure",
"question": "Are users told which assumptions support the graph?",
"artifact": "Graph with assumptions and unresolved edges.",
},
]
)
workflow| step | question | artifact | |
|---|---|---|---|
| 0 | Define the variable dictionary | What does each variable mean, when is it measu... | Measurement and timing table. |
| 1 | Encode hard constraints | Which directions are impossible because of tim... | Allowed/forbidden edge list. |
| 2 | Choose an assumption family | Are we relying on conditional independences, s... | Discovery assumptions memo. |
| 3 | Run stability checks | Does the graph change under resampling, thresh... | Edge stability table. |
| 4 | Audit caveats | Could hidden confounders, selection, measureme... | Threat model. |
| 5 | Prioritize interventions | Which uncertain edge would change the decision... | Experiment backlog. |
| 6 | Report as candidate structure | Are users told which assumptions support the g... | Graph with assumptions and unresolved edges. |
16. Decision Memo Example
Here is a concise memo for using causal discovery in an industry analytics setting.
memo = '''
### Causal Discovery Memo
**Decision context.** We want to understand which customer-health signals should be targeted to improve renewal.
**Use of discovery.** Causal discovery will be used to generate candidate causal graphs and prioritize experiments. It will not be treated as proof of causality.
**Data.** Account-month panel collapsed to pre-renewal windows. Variables include usage, support tickets, product adoption, customer-success outreach, contract value, and renewal.
**Hard constraints.**
- Renewal cannot cause pre-period usage.
- Customer-success outreach can affect later product adoption but not earlier product adoption.
- Contract value is measured before the renewal window.
- Post-renewal variables are excluded.
**Main caveats.**
- Account health is partly latent and may confound usage, outreach, and renewal.
- Support tickets are selected because only some customers contact support.
- Customer-success managers may target accounts based on unrecorded qualitative information.
- Panel dynamics mean lag choices matter.
**Analysis plan.**
- Run discovery with temporal constraints and bootstrap edge stability.
- Compare constraint-based and score-based results.
- Mark edges that are stable, unstable, or forbidden by domain logic.
- Use the graph to identify two candidate interventions for randomized testing.
**Reporting rule.** Present the graph as a hypothesis map with assumptions, not as a final causal model.
'''.strip()
display(Markdown(memo))Causal Discovery Memo
Decision context. We want to understand which customer-health signals should be targeted to improve renewal.
Use of discovery. Causal discovery will be used to generate candidate causal graphs and prioritize experiments. It will not be treated as proof of causality.
Data. Account-month panel collapsed to pre-renewal windows. Variables include usage, support tickets, product adoption, customer-success outreach, contract value, and renewal.
Hard constraints.
- Renewal cannot cause pre-period usage.
- Customer-success outreach can affect later product adoption but not earlier product adoption.
- Contract value is measured before the renewal window.
- Post-renewal variables are excluded.
Main caveats.
- Account health is partly latent and may confound usage, outreach, and renewal.
- Support tickets are selected because only some customers contact support.
- Customer-success managers may target accounts based on unrecorded qualitative information.
- Panel dynamics mean lag choices matter.
Analysis plan.
- Run discovery with temporal constraints and bootstrap edge stability.
- Compare constraint-based and score-based results.
- Mark edges that are stable, unstable, or forbidden by domain logic.
- Use the graph to identify two candidate interventions for randomized testing.
Reporting rule. Present the graph as a hypothesis map with assumptions, not as a final causal model.
17. Common Failure Modes
- Treating a discovered graph as a discovered truth.
- Forgetting that many directions are Markov equivalent.
- Running discovery on variables measured after treatment.
- Ignoring latent confounders.
- Ignoring selection into the dataset.
- Using a graph with unstable edges for high-stakes decisions.
- Mixing time scales, such as daily causes and monthly outcomes, without a timing model.
- Letting the algorithm choose between directions that domain knowledge already rules out.
- Evaluating discovery by visual plausibility rather than predictive, interventional, or stability checks.
- Skipping the next step: experiment, quasi-experiment, or validation.
18. Exercises
- In the PC skeleton example, change the significance threshold from \(0.001\) to \(0.05\). Which edges change?
- Add a weak edge \(C \rightarrow E\) to the friendly DAG simulation. How much data is needed to recover it reliably?
- In the latent confounding example, include \(U\) in the observed dataset. How does the recovered graph change?
- In the selection example, change the selection threshold from the 82nd percentile to the 95th percentile. What happens to the induced correlation?
- In the functional-model example, replace Laplace \(X\) with Gaussian \(X\). Does the residual-dependence asymmetry become weaker?
- Choose five variables from a real project. Write hard temporal constraints before running any algorithm.
- Design an experiment that would test one uncertain edge from a discovered graph.
19. Key Takeaways
- Causal discovery can suggest causal structure, but only under assumptions.
- Conditional independence can identify skeletons and some colliders, but often not unique DAGs.
- Markov equivalence means many directions are not learnable from observational independences alone.
- Latent confounding, selection bias, measurement error, feedback, and time aggregation can mislead discovery.
- Score-based and functional-model methods add different assumptions; they do not remove the need for assumptions.
- Stability checks and domain constraints are essential in applied discovery.
- In industry, the best use of causal discovery is to prioritize better causal designs, not to replace them.
References
Chickering, D. M. (2002). Learning equivalence classes of Bayesian-network structures. Journal of Machine Learning Research, 2, 445-498. https://www.jmlr.org/papers/v2/chickering02a.html
Pearl, J. (2009). Causality: Models, reasoning, and inference (2nd ed.). Cambridge University Press.
Peters, J., Janzing, D., & Scholkopf, B. (2017). Elements of causal inference: Foundations and learning algorithms. MIT Press. https://mitpress.mit.edu/9780262037310/elements-of-causal-inference/
Runge, J. (2018). Causal network reconstruction from time series: From theoretical assumptions to practical estimation. Chaos, 28(7), 075310. https://doi.org/10.1063/1.5025050
Shimizu, S., Hoyer, P. O., Hyvarinen, A., & Kerminen, A. (2006). A linear non-Gaussian acyclic model for causal discovery. Journal of Machine Learning Research, 7, 2003-2030. https://www.jmlr.org/papers/v7/shimizu06a.html
Spirtes, P., Glymour, C., & Scheines, R. (2001). Causation, prediction, and search (2nd ed.). MIT Press. https://doi.org/10.7551/mitpress/1754.001.0001
Zheng, X., Aragam, B., Ravikumar, P., & Xing, E. P. (2018). DAGs with NO TEARS: Continuous optimization for structure learning. arXiv. https://arxiv.org/abs/1803.01422